Machine learning in biological research: key algorithms, applications, and future directions | BMC Biology

Ordinary least squares regression
Overview
Ordinary least squares (OLS) is a statistical method that is used to estimate the parameters of a linear regression model [32]. OLS is sometimes also called a “best-fit line” (Fig. 4). This approach focuses on minimizing the sum of the squares of the residuals, which reflects differences between the observed values in the dataset and the values predicted by the model. In linear regression, the relationship between the dependent variable, \({y}_{i}\), and a set of independent variables, matrix \({x}_{i}\), is typically expressed as \({y}_{i}= \alpha + {\beta x}_{i}\). The coefficients \(\beta\) represent the parameters of the regression and summarize the influence of each input feature on the dependent variable. The term \(\alpha\) is the intercept and captures the baseline value of \({y}_{i}\) when all \({x}_{i}\) values are zero. The sum of squared residuals, which is explicitly the target of OLS, is given by: \({\sum }_{i=1}^{n}{\left({y}_{i}-\alpha -{\beta x}_{i}\right)}^{2}\). The least squares approach chooses \(\alpha\) and \(\beta\) to minimize the residual sum of squares. Note that usage of the squared error is an analytical convenience but can over-emphasize outlier data points. Using some calculus, one can show that the minimizing values of \(\alpha\) and \(\beta\) are:
$$\beta=\frac{\sum_{i=1}^n\left(x_i-\overline x\right)\left(y_i-\overline y\right)}{\sum_{i=1}^n\left(x_i-\overline x\right)^2}$$
$$\alpha = \overline{y } – \beta \overline{x }$$
where \(\overline x\) refers to the arithmetic mean of a variable across the data set.
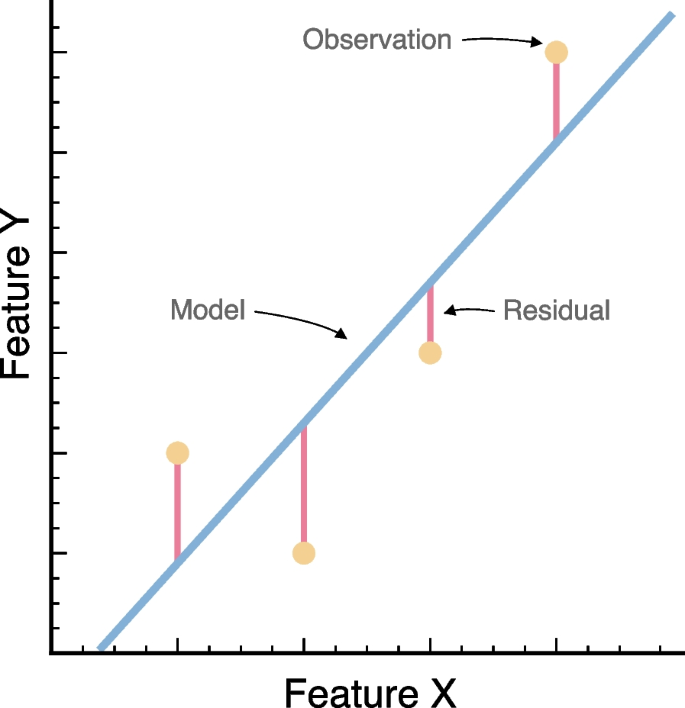
Schematic representation of a linear regression model. We summarize the scatter for a hypothetical dataset (yellow circles), a given best-fitting model (blue line), the associated residual values (vertical red lines), in accordance to a response variable and a single predictor
OLS works best when its underlying assumptions are followed, but there exist extensions for various situations. For example, by changing the squared error to an absolute error or even a median error, we can reduce the impact of outliers. Alternatively, if prior knowledge is available about the expected distribution of parameters, Bayesian regressions could provide a viable alternative to frequentist frameworks. Defining prior distributions on parameters is a form of “regularization,” which typically helps models avoid overfitting and generalize better [32]. Likewise, if the dependent variable is a discrete class, one can modify OLS into a similar model such as logistic regression. Having been deployed in the sciences for decades, there are a plethora of OLS variants for many specific situations.
Some of the major advantages of OLS relate to its flexibility, interpretability, speed, and explanatory power. Specifically, because of the expected linear relationship between the response and independent variables, one can immediately infer the effect of changing a variable value on the prediction. Further, the underlying statistics enable calculating confidence intervals both on the predictions themselves as well as the parameter values (e.g., often this criteria is used to determine inclusion of an independent variable in a model). A key approach for estimating uncertainty in parameter estimates is bootstrapping. Bootstrapping resamples the given data with replacement to create a new sample dataset of the same size. Then, parameters are re-estimated using the sample and are compared to the original parameter estimates by creating a distribution of a desired statistic (e.g., mean, median, confidence interval) for the target parameter. Finally, by requiring only elemental linear algebra, OLS is deterministic and fast. OLS often serves as a baseline against which other methods must compare.
Usage in biological research
Below, we outline two recent papers that explicitly use OLS to address questions on the intersection between ML and biology. First, Smith et al. [33] use a multiple linear regression, under a Bayesian framework (e.g., including prior distribution on regression parameters), to model the similarity between ecoregions as predicted by their geographical distance and environmental conditions. Ecoregions are large cohesive areas of land or water that are often described in terms of species assemblages, their ecological dynamics, or environmental conditions. In their paper, Smith et al. [33] use the Jaccard dissimilarity index (log-transformed) to capture the differences between ecoregions. This index is particularly used as the response variable in the examined models. Smith et al. [33] tests whether distance between ecoregions is explained by either (1) abiotic or (2) biotic factors. For their abiotic hypotheses, the independent variables were distance between regions, their mean homogeneity score, and principal components of environmental variables. For their biotic hypotheses, the independent variables were distance between regions, their mean homogeneity score, and either feeding guild or body size of terrestrial vertebrate taxa. Analyses also included the squared terms of predictors in the different models to account for a possible nonlinear relationship between their environmental predictors and distance between ecoregions. Analyses were conducted in Python using the “PyMC3” package [34]. Modifications to basic OLS include the Bayesian nature of the analysis (although uninformative priors were used). Significance of parameters was defined based on whether the relevant 95% credible intervals included 0, as is typical in statistical testing [32].
The second paper reviewed for OLS is Tao et al. [35]. In this study, the authors used linear regression models to compare estimates of phylogenetic divergence times between taxa as estimated by simple or complex models of molecular evolution. Complex models, represented in this study using GTR + Γ (general time-reversible), incorporate variable rates of nucleotide substitution. Simple models assume equal substitution rates and base frequencies. Simple and complex models were used to estimate divergence times across plant and animal clades. The explicit focus of the analyses was on node ages (i.e., branching times). Linear regression models were used to estimate the congruence between complex and simple models in terms of node age estimates. Time estimates were normalized by the sum of all node ages within each data set. The authors expected a linear pattern with low dispersion of points between the response and predictor, indicated by a slope close to 1 and high R2 values (e.g., slope = 0.95, R2 = 0.99), as a sign of high agreement between complex and simple models. We highlight that while linear regression was not the primary focus of the paper, it was particularly used to illustrate the similarity in divergence time estimates between complex models with many parameters, and simpler models, which are less computationally intensive, in the context of phylogenomic datasets.
Implementing linear regression models
Linear regression models can be fit in a number of different libraries implemented across multiple programming languages. We primarily focus on those in R or Python. Regression models can be fit using the ‘stats’ package [36]. A simple linear regression can be fit with the lm() function as indicated below. The training_data object is a table (e.g., data.frame) that includes a response variable (column y) and predictors (rest of the columns). The test_data object has the same structure of columns as training_data but was generated by splitting the full dataset into train (e.g., training_data, 70% of observations) and test sets (e.g., training_data, 30% of observations).
# Fit the model with training data
ols.model <- lm(formula = y ~ ., data = training_data)
# Make predictions with the test data
preds <- predict(ols.model, newdata=test_data)
To run an Ordinary Least Squares (OLS) regression using Python, one can use the ‘statsmodels’ library [37] and assuming that training_data and test_data are pandas DataFrames:
import statsmodels.api as sm
X = training_data[features_list]
y = training_data[target_feature]
# Fit the model with training data
# Make predictions with the test data
predictions = results.predict(test_data)
Support vector machines
Overview
Support vector machines (SVMs) are a set of supervised learning methods that are used in applications such as image classification, text classification, and various of bioinformatics routines. SVMs are often used for classification but can also be adapted for regression tasks. Similarly, although SVMs are generally fit for supervised learning, variations of SVMs can also be used under an unsupervised framework (e.g., one-class SVM). Before the 1980s, almost all learning methods learned linear decision surfaces, and the amount of samples in theoretical statistical studies was assumed to be large or infinite to simplify mathematical analyses. However, the size of empirical datasets is usually limited, and the relationships between features are almost never linear. In 1995, Vladimir N. Vapnik developed a novel approach and showed that SVMs work well with nonlinear and high-dimensional datasets at pattern recognition routines [5]. Based on the concept of similarity, SVMs use nonlinear “kernel” functions to transform the data to a higher dimension, enabling linear separation by finding optimal boundaries (i.e., hyperplanes) that form the best partition (i.e., decision boundary) between (1) classes and (2) support vectors or the data points that lie closest to the decision surface (or hyperplane) to maximize the margins between the classes (Fig. 5). SVMs are flexible in defining similarity measures and often generalize well to new data. With the advantage of global optimization and strong adaptability, SVMs have wide applications in areas like protein classification, and computer vision, among others.
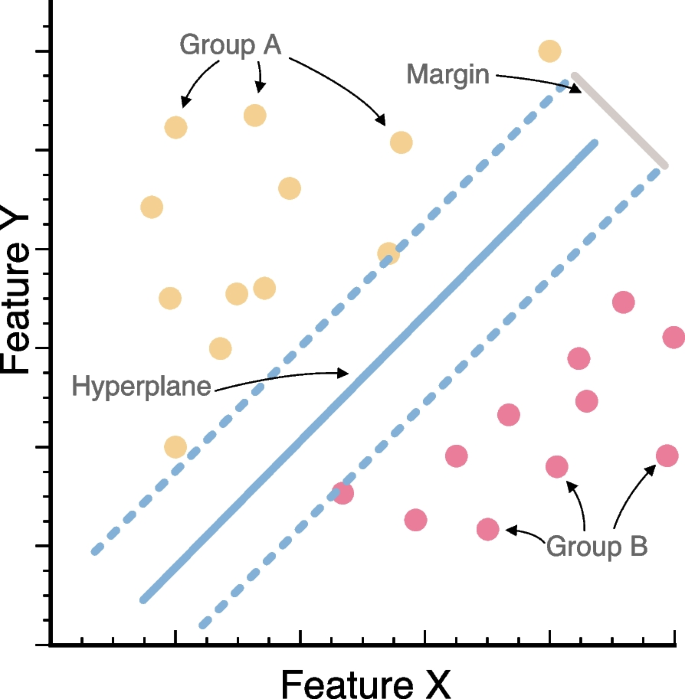
Visual representation of support vector machines (SVMs) and its key elements. We present the relationships between two features in accordance to two different classes (group A in yellow, and group B in red). We show the hyperplane dividing the two groups, as well as the margin summarizing the overall division between classes
SVMs are primarily focused on determining the hyperplane that optimally divides data into particular classes based on the maximum margin [5, 38]. The hyperplane is defined such that the minimum distance between data points in different groups (i.e., support vectors) is maximized. In the case of pairwise data, \(\left(x_i,y_i\right)\), where \(\left({x}_{i} \in {R}^{n}\right)\) are the feature vectors and \(\left({y}_{i} \in \left\{1, -1\right\}\right)\) are the class labels, SVMs are focused on solving the following optimization problem:
$$\left[\min_{w,b}\frac12\left.\Arrowvert w^2+C\sum\nolimits_{i=1}^m\xi_i\right]\right.$$
subject to the constraints \(\left(y_i\left(w\cdot x_i+b\right)\geq1-\xi_i\right)\) for every \(i\). Note that \(\left({\xi }_{i} \ge 0\right)\) are slack variables that allow misclassification of either challenging or noisy points. Similarly, \(C\) is a regularization parameter that enables the control of the trade-off associated with achieving a high margin while reducing training error. The minimization itself, however, typically requires an iterative approximation as the non-linear kernel often precludes an analytical solution.
Kernels are the source of SVMs’ intrinsic flexibility [39]. Kernels allow for operations in the input space to be equivalent to operations in a higher dimensional feature space [40]. These operations based on kernels occur implicitly without the need of computing coordinates in the novel space [41]. For instance, assume that two populations of a given species inhabit distinct elevations and this is the key feature distinguishing them. However, elevation was not included as a feature in the dataset. Under SVMs, the use of a certain kernel on the features that were effectively collected in the dataset (e.g., latitude and temperature) could result in the indirect inclusion of elevation as a result of the expanded multivariate space (e.g., a proxy of elevation).
Mathematically, an SVM kernel function is the dot product of two vectors of higher dimensional space. Commonly used kernels functions polynomial kernel \(k\left(x_i,x_j\right)=\left(\gamma x_i\cdot x_j+r\right)^d\), radial basis function (RBF) kernel \(k\left(x_i,x_j\right)=\exp\left(-\gamma\Arrowvert x_i-x_j\Arrowvert^2\right)\), and sigmoid kernel \(k\left(x_i,x_j\right)=\tanh\left(\gamma x_i\cdot x_j+r\right)\), where \(\gamma\), \(r\), and \(d\) are parameters that can be adjusted based on the data set.
Mathematically, an SVM kernel function generally involves the dot product of one data point with another, \(\langle x_i,x_j\rangle=\sum\nolimits_kx_{ik}x_{jk}\) where \(k\) indexes a given feature in the vector (e.g., temperature, latitude). These intermediate dot products can then feed into much more general non-linear functions such as the linear kernel or sigmoid kernel. The specific choice of kernel is beyond the scope of this review and is considered part of the larger model learning procedure (but see [32] for a detailed discussion). Because of this capability for handling non-linearity, SVMs excel at domains where it is possible to draw a continuous “boundary” between data points of different classes. The nature of the kernel determines the shape capabilities of this boundary (e.g., a linear kernel will have boundaries which are linear in the independent variables).
When fitting SVMs, practitioners typically focus on fitting three critical parameters to optimize their models. First, the choice of the kernel type determines the transformation space of the input data. Each kernel type is suited for different types of data. For instance, the linear kernel is preferred for data that are linearly separable in the input space. The RBF kernel can handle more complex, nonlinear relationships. Second is tuning the regularization parameters, particularly the penalty parameter \(C\) and the kernel-specific parameter \(\gamma\). These two parameters are essential for preventing overfitting and ensuring that the model generalizes well to new data. \(C\) controls the trade-off between achieving a low error on the training data and minimizing the model complexity for better generalization. The \(\gamma\) parameter defines how far the influence of a single training example reaches: low values indicate “far” and high values indicate “close.” Third, defining the optimal value for the margin (i.e., the decision boundary) is crucial. A larger margin can increase the generalizability of the classifier. However, if the margin is set too wide, it might lead to misclassification of the training data, especially if the data are noisy or not well-separated.
Usage in biological research
We selected two case studies using SVMs. One paper focuses on detecting leaf disease using images [42] and the second on inferring taxonomic information of hosts based on viral genomes [43]. First, Das et al. [42] implemented a classifier to identify healthy and unhealthy tomato plants based on photos of their leaves. The authors focused on developing classifiers that could help improve the agricultural sector in India, ultimately enhancing the living standards of the rural population. For this study, the authors collected images from an existing database containing images of healthy and unhealthy tomato leaves (n = 14,000). They conducted image preprocessing steps and masking, including resizing and conversion to grayscale for further marking of target pixels. Color was extracted from RGB channels based on the masked images. These features (e.g., RGB channels, texture, contours) were then used to train and test random forest, logistic regression, and SVMs based on healthy and unhealthy classes. The training phase of the model was conducted on 60% of the images. The testing set, used for assessing model performance, comprised the remaining 40% of the observations. Das et al. [42] recovered a 25–30% higher accuracy for SVMs compared to random forest and logistic regression models. Based on these results, Das et al. [42] supported the deployment of SVM models for real-life applications in automatic disease detection at early stages.
Second, Young et al. [44] aimed to increase the available information on hosts for newly described viral genomes. The majority of newly discovered viruses lack taxonomic information for host species. The goal of this study was to identify genomic features of the virus that could be used to accurately predict the taxonomic information of the host. The key challenge was representing viral genomes in a format that made discriminative information available for ML procedures. For this study, sequences were retrieved from Virus-Host Database (VHDB) and RefSeq. Genomes were summarized as nucleotide sequences, amino acid sequences, physicochemical properties, and predicted PFam domains. From these representations, k-mer or domain extraction procedures were conducted to obtain a feature matrix. SVMs were trained on 80% of the data, with testing conducted on the remaining portion of the dataset (75% vs. 25% in alternative analyses). Phylogenetic information was accounted for in the analyses using a “holdout” method including an average nucleotide identity filter. The SVM used a linear kernel, and performance was evaluated based on not just overall accuracy, but also using receiver operating characteristic (ROC) curves (equivalent to precision-recall curves and other approaches that simultaneously account for both false positive and false negative errors). The authors also combined different types of features derived from the same viral genomes and assessed their ability to predict host information. Based on their SVMs, Young et al. [44] found that all the analyzed feature sets were predictive of host taxonomy. However, combining feature sets had the potential to improve predictive accuracy further.
Implementation of SVMs
SVMs are implemented in a range of libraries both in Python and R. In R, the e1071 package [45], a wrapper of the LIBSVM C++ library, is standard for implementing SVMs.
model <- svm(y ~ ., data = training_data, type=”eps-regression”, kernel=”linear”)
predictions <- predict(model, newdata = test_data)
model <- svm(formula = class ~ ., data = training_data, type=”C-classification”, kernel = ‘linear’)
predictions <- predict(model, newdata = test_data)
Alternatively, excellent implementations of SVMs can be found in tidymodels, a package that offers a streamlined and modern framework for ML modeling within R [46]. The package includes functions such as svm_rbf() and svm_linear() that facilitate the application of SVMs with different kernel types in both regression and classification tasks.
In Python, the primary library for implementing SVMs is scikit-learn [47]. This versatile library implements functions to fit and analyze SVMs including SVC (Support Vector Classification), SVR (Support Vector Regression), and LinearSVC, an implementation that supports linear and non-linear SVMs. Additionally, the BioPython toolkit and related libraries offers closely integrated SVM-related implementations [48];
from sklearn import svm
model = svm.SVR(kernel=”linear”)
# Fit the model with training data
# Make predictions with the test data
predictions = model.predict(test_data)
model = svm.SVC(kernel=”linear”)
# Fit the model with training data
# Make predictions with the test data
predictions = model.predict(test_data)
Random forest
Overview
Random forest (RF) is a machine learning technique that has gained widespread popularity among researchers and practitioners due to its versatility and effectiveness, particularly in prediction tasks [49, 50]. This approach builds on an ensemble of decision trees that, besides predictive and inferential tasks, enable feature selection procedures as part of analyses and explicitly model interactions between variables [49, 51,52,53]. RF belongs to the ensemble learning family, a framework that combines multiple individual models to improve overall predictive performance. The “forest” in random forest is composed of decision trees (Fig. 6). A decision tree resembles a flowchart structure where each internal node represents a threshold or definition based on a particular feature. Branches represent decision rules and each leaf node represents an outcome. Decision trees are simple and easily interpretable yet effective models for classification and regression procedures [32].
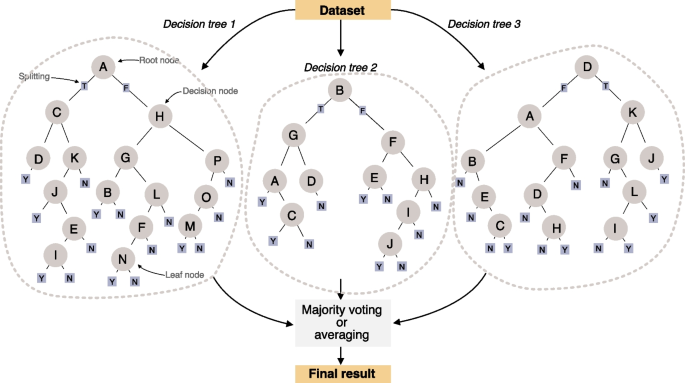
Illustration of a random forest algorithm based on three decision trees and a number of features used to train the model (letters in circles). We show different decision trees that are trained independently from an initial dataset. Results across decision trees are summarized using either majority voting or averaging
There are at least six aspects that are critical to understand the structure, fit, and performance of RF algorithms. First, RF employs a sampling technique called bagging. This approach involves training each decision tree on a random subset of the training data (sampled with replacement, so a data point can occur multiple times for the same tree) that ultimately reduces overfitting by introducing diversity among trees. Second, each decision tree in the RF is constructed using a subset of features selected randomly at each node. This randomness ensures that trees are less correlated with each other, leading to a more robust model. Third, one of the hyperparameters of RF is the number of trees in the forest. Typically, increasing the number of trees improves performance while increasing computational cost. Finding the optimal number of trees often involves cross-validation techniques (i.e., trying many different values on subsets of the data while scoring on held-back data). Fourth, RF provides a measure of feature importance, which indicates the contribution of each feature in predicting the target variable. This information can be useful for feature selection and understanding the underlying data. Fifth, training each decision tree in RF is independent of the others, making it highly parallelizable. Many implementations of RF leverage parallel computing to speed up the training process, especially when dealing with large datasets. Sixth and finally, RF has several hyperparameters such as the number of features to consider at each split, maximum depth of the trees, minimum samples per leaf, among others to offer additional flexibility. Grid search or randomized search techniques can be used to find the optimal combination of hyperparameters [32]. These key aspects all contribute to the ongoing effectiveness and popularity of RF.
Note that there are a variety of tree-based ensembles that are structurally similar to random forests. For instance, we will describe gradient boosted trees later in this review. Bayesian additive regression trees are also a popular approach that also fall within the tree-based ensemble framework [54]. Each method, however, has a different training procedure. Just as Bayesian linear regression has the same structure as OLS, tree ensembles can come in many forms with different trade-offs.
Usage in biological research
We explore the use of random forest in two case studies. First, Fabris et al. [52] used random forest to discern loci underlying both discrete and quantitative traits, particularly when studying wild or non-model organisms. RF is becoming increasingly used in ecological and population genetics because, unlike traditional methods, it can efficiently analyze thousands of loci simultaneously and account for non-linear interactions. The authors described how to prepare data for RF, including initial data exploration, the identification of important features, and possible confounding factors. They then provided guidance on the initiation of RF and the optimization of the algorithm parameters for classification and regression. Finally, they summarize methods for interpreting the results of RF and identifying trait-associated or predictor loci. Second, Brieuc et al. [51] focused on looking at how RF can be used effectively in studies focused on genotype–phenotype associations, particularly in non-model organisms. This study, structured as an introductory guide to the intersection between RF and ecological and evolutionary genomics, discusses fundamental approaches to carefully fit, analyze, assess performance, and understand results of RF-based approaches.
Implementation of random forest models
We present implementations of random forests using Python and R. In R, random forests can be fit with a given dataset using the randomForest package [55] as follows:
# Initialize the random forest model
rf_model <- randomForest(x = X_train, y = y_train, ntree = 100, importance = TRUE, classwt = “balanced”)
# Make predictions on the test data
rf_pred <- predict(rf_model, X_test)
Random forests can also be implemented in Python using the scikit-learn library [47]:
from sklearn.ensemble import RandomForestClassifier
# Initialize the Random Forest model
rf_model = RandomForestClassifier(max_depth=2, random_state=0)
rf_model.fit(X_train, y_train)
# Make predictions on the test data
rf_pred = rf_model.predict(X_test)
Note that where SVMs have a few choices of kernel function and possibly one or two other parameters, RF has a plethora of so-called hyperparameters. One can vary the number of trees, features considered in splitting, the required purity for a node to stop being split, and more. Implementations of RF generally expose these parameters at model initialization, implying that trying different options via cross-validation or similar techniques is recommended. Methods like grid search, random search, or Bayesian optimization are common techniques for systematically optimizing these hyperparameters. Specifically, grid search exhaustively searches through a specified range of parameter values, while random search randomly samples from a distribution of parameter values. Bayesian optimization uses probabilistic models to focus the search on promising regions of the parameter space. Cross-validation is typically used to evaluate the performance of different parameter configurations and select the best one. Additionally, Out-Of-Bag (OOB) performance metrics are also commonly used in RF. OOB refers to the data points excluded from a bootstrap sample when training individual trees in an ensemble model. These data points are used as a built-in validation set to estimate the predictive power of each tree within the forest [11, 33].
Gradient boosting
Overview
Gradient boosted models (GBMs) can be understood by extending our prior explanation of random forest. Where random forest [49] creates an ensemble of trees through bagging, gradient boosting develops each component model (i.e., individual decision tree) of the ensemble one after the other [32]. This iterative procedure is generally called boosting (Fig. 7). Specifically, let \({f}_{m-1}\left({x}_{i}\right)\) be the boosted model’s prediction after \(m-1\) components have been added. Under this boosting, we seek the next iteration,\(f_m\left(x_i\right)=f_{m-1}\left(x_i\right)+\Gamma_mg_m\left(x_i\right)\) (i.e., one has generated two successive models via GBM and seek to refine it by adding a third component to the ensemble). For example, one could fix \({\Gamma }_{m}=1\) and fit \({g}_{m}\) to minimize the residual loss, \(L\left({y}_{i}-{f}_{m-1}\left({x}_{i}\right), {g}_{m}\left({x}_{i}\right)\right)\). That is, each new component attempts to correct errors of the previous model. The way to determine \({g}_{m}\) and \({\gamma }_{m}\) depends on the exact nature of the boosting. One subtype of boosting is called gradient boosted models [56] or GBMs. This approach fits \({g}_{m}\) to minimize the loss on the negative gradient,\(-\frac{\partial L\left({y}_{i}\right)}{{\partial f}_{m-1}}\). Then one finds the weight, \({\Gamma }_{m}\) to minimize the overall loss, \(L\left({y}_{i}, {f}_{m-1}\left({x}_{i}\right)+{\Gamma }_{m}{g}_{m}\left({x}_{i}\right)\right)\). The gradient helps direct the next model more carefully than generic boosting.
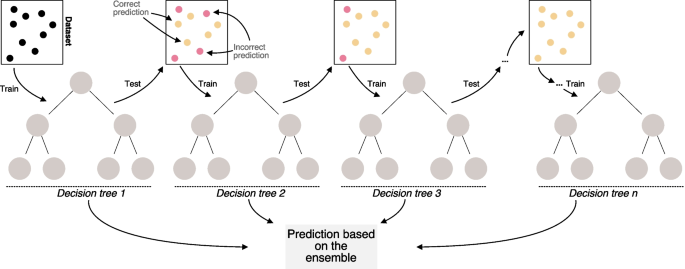
Diagram showing the temporal patterns in training for Gradient Boosting Trees. We show an initial dataset used for training of a decision tree. Subsequent iterations of the boosting sequence are focused on enhancing model performance by reducing error rates. We show correct model-based predictions in yellow. Red circles summarize model errors. At the end, model-based predictions are performed on the tree ensemble
The exact capabilities of GBMs depend strongly on the underlying model type within the ensemble. For example, gradient boosted trees (GBTs) are structurally identical to random forests, and they are often applicable to similar problems. Because GBMs use a gradient, however, they can take advantage of a continuous loss function to speed-up model convergence. Conversely, problems with discontinuous losses may be less suitable for GBMs.
Usage in biological research
We selected two case studies that summarize the use of gradient boosting in the context of biological research. First, Zhang et al. [57] built a predictive model for bioluminscent proteins (BLPs) via sequence-derived features for identification. BLPs are valuable for both industry and research. In this study, the authors used XGBoost (eXtreme Gradient Boosting), an ensemble learning algorithm based on gradient boosted trees [58]. XGBoost is well known for its highly flexible and scalable tree structure enhancement model and its reduction of computational time and memory for training large-scale data. All of these features were used to specifically improve on methods and tools previously used for the prediction of BLPs. First, a previously constructed comprehensive dataset consisting of BLP sequences and non-BLP sequences composed from bacteria, eukaroyte and archaea, was collected from UniProt to be used as training and predictive data. In order to avoid homology bias, the data was first cleaned by using BLASTClust [59]. The variety of features which make these sequences identifiable and which was characterized by previous work, was further encoded via various methods (i.e., natural vector, composition/transition/distribution, g-gap dipeptide composition, and pseudo amino acid composition), mainly so that the data could be processed through ML algorithms like XGBoost. The dataset was then used to develop the prediction model by finding the highest area under the curve (AUC) values (again, a measure of overall accuracy and the trade-off against false alarms) correlated to specific encoded features for which each of the species in the set could be optimized for prediction. Performance was then measured by testing the data using different combinations of encoding features along with XGBoost’s internal statistical analyses for cross-validation. Results indicate strong predictions for species-specific trained data sets and overall more accurate results when compared to other predictive algorithms (e.g., decision trees, random forests, and AdaBoost).
Second, Yaşar et al. [43] used multiple ML predictive algorithms (deep neural network, random forest, and GBT) to classify three COVID-19 positive patient groups (mild, severe, and critical). This study also aimed to generate a control group by using blood protein profiling as predictive indicators. The team obtained a dataset that included age, gender, and 368 proteins obtained from blood protein profiling. The number of people in each severity group and control group varied. In order for the data to be successfully used by the algorithms, it first had to be standardized in multiple ways to account for inequalities (e.g., missing values, unbalanced sample size). After the data was cleaned, the ML algorithms were trained to make predictions about disease severity. The GBT algorithm was worked in as noted,a prediction function was iteratively refined. Residuals computed from the difference between predictions and actual data subsequently informed the next target in GBT, creating new and more accurate residuals which were used as training data as further iterations occurred. Using ML predictive algorithms, the authors identified 10 proteins associated with COVID-19 severity that could be used as bio-markers, with two of them (IL6 and LILRB4) supporting results of previous proteomic-based work. GBT achieved the best prediction of disease severity based on available proteins compared to the rest of the tested algorithms according to the results of various metrics (i.e., accuracy, sensitivity, specificity, precision, classification error, and others).
Implementation of Gradient Boosting
Libraries to implement GBMs exist in many programming languages. For this review, we offer two code snippets, one in R and one in Python, to demonstrate one way to deploy this method in a simple dataset, split into train and test sets. First, GBMs can be fit in R using the gbm package [60] as follows,
“`r library(gbm) ## Build a predictive model for regression or classification # There are several tuning parameters in the gbm package that this example code leaves out for simplicity. Users can specify the number of trees, interaction depth, and cross-validation folds, among other parameters, to tune the model. # Train the model model<- gbm(y ~ ., data = training_data) # Using the model, make predictions on the test data predictions<- predict(model, test_data)“`
Alternatively the xgboost library in Python can be used to fit a Gradient Boosted Trees model with XGBoost [58]
“`python import xgboost as xgb # For regression # Initialize the model reg = xgb.XGBRegressor(n_estimators=10) # Train the model reg.fit(X_train, y_train) # Make predictions on the test data predictions = reg.predict(X_test) #For classification # Initialize the model clf = xgb.XGBClassifier(n_estimators=10) # Train the model clf.fit(X_train, y_train) # Make predictions on the test data predictions = clf.predict(X_test)“`
Challenges of ML-based inference in biology
Despite the flexibility and potential of machine learning models in biological research demonstrated through the examined case studies, ML as an analytical framework still suffers from various constraints that limit its performance and widespread use within disciplines. In this section, we briefly review the major pitfalls associated with using a machine learning framework to address questions of biological relevance. We focus on the models examined in this paper, detailing the limitations of machine learning in biology and examining the potential for future advancements. It is essential to critically evaluate limitations to best leverage machine learning in biological research. Furthermore, the applicability and reliability of these models in the field can directly be understood from exploring innovative solutions to these challenges.
A common challenge when applying machine learning as an analytical framework to answer biological questions relates to selecting the most effective technical implementation of a given algorithm (see also [61]). For instance, although this paper presents only a single SVM framework, there exists a wide range of alternative approaches for analyzing SVMs (e.g., svmSomatic, GraDe-SVM, MD-SVM). The same situation applies to GBMs (e.g., LightGBM, XGBoost, CatBoost), linear regressions (e.g., ridge regression, lasso regression, Bayesian linear regression), random forests (e.g., extra trees, oblique random forests, rotation trees), and most—if not all—algorithms within a machine learning framework. At this point, users will likely face several important questions. First, which of the available alternatives should be used? Second, do different implementations affect the results? We encourage less experienced users to evaluate model performance using the simplest version of the implementation before exploring other alternatives (e.g., [62, 63]). In some cases, explicit constraints or specific data structures justify the use of question-specific models (e.g., TF-Boosted Trees, a TensorFlow implementation for structured data problems). In such cases, users should prioritize more specialized approaches in accordance with their knowledge of the data, the question, and the algorithm. Trying multiple algorithms is an option—and often common practice—but more importantly, users should critically assess why fitting a particular method is advantageous as an a priori step before examining its performance.
Another critical aspect to machine learning, closely linked to both model fitting and performance assessment, is data visualization (e.g., visual analytics; [64,65,66]). Despite being oftentimes overlooked in favor of model-focused paradigms when machine learning is introduced, data visualizations are integral to understanding both the process and results of machine learning. In many cases, model performance can be evaluated directly by visualizing relevant features, particularly when guided by domain knowledge [67]. For instance, in classification tasks, visualizations such as logistic regression nomograms can provide a clear understanding of how feature values influence a model’s predictions. These visual representations allow researchers to distinguish patterns that may not be immediately apparent in raw data or numerical outputs. Furthermore, assessing model performance or identifying violations of assumptions often involve creating plots such as residual plots (e.g., regression) or ROC curves (e.g., classification, [62]). These types of visual tools can help reveal the accuracy, precision, and limitations of a model. Interpretability is inherently tied to data visualization, as it transforms complex, often abstract model behavior into a format that can be easily understood. We highlight that the true success of a machine learning model lies not just on its ability to predict a given outcome accurately, but in its capacity to explain that prediction in an accessible manner [68, 69]. Models may capture patterns—some visible, other invisible—that, when visualized appropriately, offer valuable insights into underlying biological phenomena [58]. Data visualizations, therefore, are not only a tool for model validation but also for conveying the nuances of model performance. Visualizing data, models, and their performance make the entire process more transparent and interpretable.
In the following paragraphs, we will discuss the intersectionality between each of the four algorithms outlined in this review, in the context of biological research. Although OLS is a basic and widely used method in machine learning, it relies on multiple assumptions that are often not met by many datasets (e.g., constant error variance and independent data points). For instance, in a dataset including phylogenetically related organisms, OLS may fail to reveal relationships present within but not across clades. Conversely, OLS might indicate relationships across clades that do not exist once phylogeny is accounted for. In such cases, phylogenetic generalized least squares (PGLS) explicitly incorporates information on relationships between terminals through a variance–covariance matrix and may be a better approach [70]. This example shows that the structure of the dataset is highly relevant for ultimately selecting the appropriate analytical tools. Even though OLS has been shown to be robust to some violations of its assumptions (e.g., [71, 72]), there should be a particular focus on reviewing the appropriateness of the method in accordance with the focal question. Although linear regression models are straightforward to implement, researchers should consider whether their aims are predictive or causal. For causal inference, careful selection of predictors is crucial. Tools such as directed acyclic graphs should be used to make explicit the hypothesized relationships between variables [73]. In some scenarios, alternative modeling approaches may better accommodate the complexities of biological data. A flexible and iterative framework of model selection and evaluation can enhance the robustness of the findings.
In the context of SVMs, preprocessing and interpretability are critical aspects of this approach. First, preprocessing of datasets analyzed using SVMs generally involves several crucial steps. Normalization is critical as SVMs are not scale-invariant, requiring features to be scaled to have zero mean and unit variance to ensure the model does not bias toward attributes with higher magnitude values. Imputation, the process of replacing missing data with an estimated values, is necessary for SVMs to be successful. Missing values are common in biological datasets. Balancing is particularly important in classification tasks where class distributions are uneven, as unbalanced datasets can lead to biased models that overpredict the majority class. Thus, developing new or systematically enhancing the existing preprocessing techniques can significantly improve the performance of SVMs in biological applications. Second, relative to simpler models, the interpretability of SVMs is generally challenging. Understanding the estimated weights is the primary focus, but the use of kernels often leads to the exploration of novel multivariate spaces, making it difficult to interpret results in the context of the biological question. Despite these challenges, we reviewed two case studies in which SVMs were successfully used to answer questions in the field mostly based on model performance (not on variable importance). Enhancing the interpretability of SVMs, either through innovative visualization approaches or explanation methods, could make these models more accessible to researchers.
Key aspects to account for when analyzing random forest models include overfitting, data requirements, model complexity, validation, and generalization. First, while RF is more robust against overfitting compared to individual decision trees, overfitting can still occur, especially with noisy or high-dimensional biological data. Careful tuning of hyperparameters such as the number of trees, maximum depth, and minimum samples per leaf is necessary to mitigate overfitting and ensure generalization to unseen data. Another approach to parameter tuning involves incorporating domain-specific knowledge into the model tuning process. Depending on the dataset used with RF, the model is more sensitive to changes in different hyperparameters. For example, Huang and Boutros [74] found that in a next-generation sequencing quality-control dataset with a low p/n ratio (variables/samples), the mtry parameter (number of variables to sample) had a significant impact on the resulting model, while the number of trees and sample size did not. In an mRNA abundance dataset with a high p/n ratio, all three parameters (mtry, number of trees, and sample size) had a significant impact on the resulting model [74]. Employing domain knowledge allows researchers to focus on tuning specific hyperparameters that have the greatest impact on the model. This external knowledge can enhance the applicability of RFs into specific datasets. Second, RF performs well with large datasets. However, biological datasets often pose unique challenges such as imbalanced class distributions, missing values, and high dimensionality. Pre-processing steps like feature selection, imputation, and data balancing are thus crucial to optimize model performance and prevent biases. Third, RF can handle complex relationships between genetic variables and phenotypic traits, but it may not capture subtle interactions or continuous relationships present in biological systems. Model interpretation and validation techniques, such as permutation importance and partial dependence plots (i.e., visualizations used to examine how model predictions perform as (i) one or multiple inputs change while (ii) the rest remains constant), help elucidate the relationships between genetic predictors and ecological or evolutionary outcomes. Fourth, assessing the performance and generalization of RF models across different populations, species, or environmental conditions is essential in biological studies. Cross-validation techniques, independent validation datasets, and robustness testing help ensure the reliability and applicability of random forest models in diverse ecological and evolutionary contexts.
Finally, GBMs have a powerful predictive performance. However, this outstanding performance comes with a set of challenges that often needs to be addressed in order to maximize their utility in biological research. First, GBMs are prone to overfitting, especially with noisy or high-dimensional biological data. Appropriately tuning parameters such as learning rate, number of trees, and tree depth is thus crucial to achieve an optimal performance across different datasets. Second, the computational complexity of GBMs can be high, often requiring significant computational resources and time. Resource-related limitations are often one of the reasons why GBMs are limited to smaller datasets or procedures of low complexity. Third, the interpretability of GBMs is generally lower compared to simpler models [75, 76]. In this context, the ensemble of decision trees can obscure the understanding of feature importance and interactions (see also RF). However, techniques like SHapley Additive exPlanations (SHAP) values or partial dependence plots are often used to provide details of the model’s decision-making process [77, 78]. SHAP values, for example, measure the contribution of each feature to individual predictions [77]. SHAP thus allows for a more granular understanding of model behavior in the context of the relevant task. Partial dependence plots, on the other hand, show the relationship between a selected feature and the predicted outcome. These plots show how changes in feature values impact model predictions. Future developments in explainable AI may enhance the interpretability of GBMs, potentially making these models more user-friendly for biological researchers. Fourth, handling missing data and imbalanced class distributions in biological datasets requires robust preprocessing strategies that ensure GBMs to produce unbiased predictions [79]. Missing data and class imbalance are, however, common in biological datasets. Lastly, rigorous cross-validation, independent test sets, and domain-specific adjustments to the model are often needed to ensure the generalization and validation of GBMs across different datasets (e.g., varying species, environmental conditions, or population structures).
Outlook: deep learning in biology
Although the main focus of this review has been on foundational machine learning and its intersection with biological fields, deep learning (DL) methods (i.e., neural networks) have been gaining strong relevance in multiple subdisciplines due to their flexibility. The basic DL framework is inspired by neuronal connections. Nodes in the graph, often referred to as neurons, are connected in layers of variable length. The first layer of neurons processes the input features and the final layer computes the output, using the previous layer as input. In its simplest form, neurons in each layer share connections to neurons in adjacent layers. The model is generally parameterized by a learning algorithm that propagates feedback to the internal weights attached to neurons from the training data through these connections. This results in a parameter-rich model capable of decomposing meaningful signals from highly complex feature sets [80].
Numerous variants of neural networks and learning models have emerged over the years. Deep learning techniques that excelled at image recognition were being applied in biological sciences going as far back as the late 1990s (e.g., [81,82,83]). Early applications were limited to segmentation of medical imagery [3], disease recognition [84], and diagnosis [85]. Classification models have demonstrated impressive accuracy on test data [86,87,88] but await refinement before adoption into widespread clinical use [89]. Other examples of early explorations can be found in drug discovery [90], virtual screening [91], and functional genomics [90, 92]. Deep learning methods have amassed greater traction from the biological science community in recent years. For instance, following rapid advances in high-throughput sequencing and the popularization of resequencing-based studies [93], deep learning techniques have made notable strides in variant detection for molecular pathologies [94, 95]. First in CASP13 in 2018 [96] and later again CASP14 in 2020 [97], Google DeepMind’s AlphaFold triumphed over competitors demonstrating resounding classification performance on protein folding patterns. Today, advancing medical technologies, the promise of personalized medicine and an abundance of DNA sequence data at the population scale have placed biological and life sciences at the forefront of explorative research in deep learning [98].
Recent breakthroughs in deep learning (e.g., transformers and large language models such as ChatGPT and Gemini) have initiated a new age of generative AI (Artificial Intelligence; [99]). Both public and commercial attention to this field of science has never been greater. There has been a cumulative increase in data center and research funding from government and private sectors [100,101,102]. Dramatically improved parallel computing infrastructure made possible by fabrication of as low as 3 nm process semiconductors [103] and high-speed interconnect between compute units [104] pave the way for greater advancements. These generative approaches show some promise, especially in their breadth of apparent knowledge, but they are prone to “hallucination,” sometimes fabricating plausible but incorrect or untested concepts as a result of their probabilistic nature and weakly curated data sets [105]. As the subfield of artificial intelligence and deep learning keeps gaining momentum, reintroduction of proven methods and the development of new algorithms tailored to the needs of biological sciences is essential.
Our review presents DL and ML as viable approaches for answering questions in biological disciplines. The choice between traditional machine learning and deep learning hinges on the nuanced trade-offs among data availability, computational resources, model interpretability, accuracy, and scalability, demanding careful consideration tailored to the specific requirements and constraints of each biological research question. Below, we present three key aspects that highlight the pragmatic difference between traditional ML and DL. First, the relevance of DL is primarily evident in applications where large semi-structured data sets are available (e.g., images of cells instead of tables). Relative to traditional ML algorithms, large DL models have demonstrated an unprecedented ability of processing language and visual data and demonstrated crude deductive capabilities [106, 107]. Second, when data is limited, traditional machine learning methods often outperform DL models, which require extensive datasets for optimal performance. The drawback of DL models is their requisite dependence on an abundance of high quality and often manually curated training data [108]. Second, the training time required for DL models can significantly exceed that of traditional methods, thus influencing decisions based on computational resources and time constraints. Recent advances in parallel computing hardware and community-accelerated software development have enabled the construction of gargantuan models comprising billions of parameters, though training speed remains an issue for specialized analyses [109]. Third, method complexity and interpretability remain critical considerations,while DL techniques often achieve superior accuracy in complex pattern recognition tasks, this accuracy frequently comes at the cost of reduced interpretability and transparency. Conversely, foundational machine learning models such as linear regressions or decision trees offer greater interpretability, making them preferable in scenarios where explainability and transparency are crucial, even if they may occasionally sacrifice a degree of predictive accuracy.
link