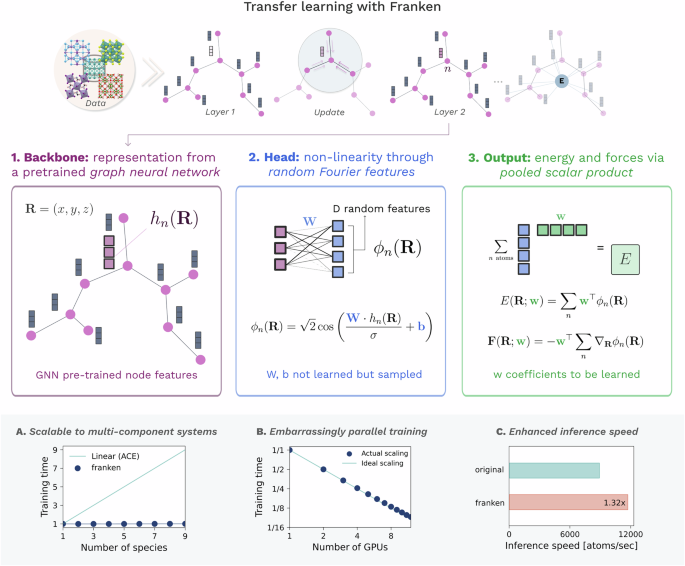
Fast and Fourier features for transfer learning of interatomic potentials
Frenkel, D. & Smit, B. Understanding molecular simulation: from algorithms to applications (Elsevier, 2023).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Google Scholar
Behler, J. Atom-centered symmetry functions for constructing high-dimensional neural network potentials. J. Chem. Phys. 134, 074106 (2011).
Google Scholar
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B Condens. Matter Mater. Phys. 87, 184115 (2013).
Google Scholar
Imbalzano, G. et al. Automatic selection of atomic fingerprints and reference configurations for machine-learning potentials. J. Chem. Phys. 148, 241730 (2018).
Google Scholar
Kývala, L. & Dellago, C. Optimizing the architecture of Behler-Parrinello neural network potentials. J. Chem. Phys 159, 094105 (2023).
Google Scholar
Schölkopf, B. & Smola, A. J. Learning with kernels: support vector machines, regularization, optimization, and beyond (MIT Press, 2002).
Shawe-Taylor, J. & Cristianini, N. Kernel methods for pattern analysis (Cambridge University Press, 2004).
Bartók, A. P., Payne, M. C., Kondor, R. & Csányi, G. Gaussian approximation potentials: The accuracy of quantum mechanics, without the electrons. Phys. Rev. Lett. 104, 136403 (2010).
Google Scholar
Drautz, R. Atomic cluster expansion for accurate and transferable interatomic potentials. Phys. Rev. B 99, 014104 (2019).
Google Scholar
Zhang, L., Han, J., Wang, H., Car, R. & E, W. Deep potential molecular dynamics: a scalable model with the accuracy of quantum mechanics. Phys. Rev. Lett. 120, 143001 (2018).
Google Scholar
Reiser, P. et al. Graph neural networks for materials science and chemistry. Commun. Mater. 3, 93 (2022).
Google Scholar
Batatia, I. et al. The design space of e (3)-equivariant atom-centred interatomic potentials. Nat. Machine Intell. 1–12 (2025).
Fu, X. et al. Forces are not enough: Benchmark and critical evaluation for machine learning force fields with molecular simulations. Transactions on Machine Learning Research (2023).
Kulichenko, M. et al. Data generation for machine learning interatomic potentials and beyond. Chem. Rev. 124, 13681–13714 (2024).
Google Scholar
Jinnouchi, R., Miwa, K., Karsai, F., Kresse, G. & Asahi, R. On-the-fly active learning of interatomic potentials for large-scale atomistic simulations. J. Phys. Chem. Lett. 11, 6946–6955 (2020).
Google Scholar
Schran, C., Brezina, K. & Marsalek, O. Committee neural network potentials control generalization errors and enable active learning. J. Chem. Phys. 153 (2020).
Bonati, L. & Parrinello, M. Silicon liquid structure and crystal nucleation from ab initio deep metadynamics. Phys. Rev. Lett. 121, 265701 (2018).
Google Scholar
Abou El Kheir, O., Bonati, L., Parrinello, M. & Bernasconi, M. Unraveling the crystallization kinetics of the Ge2Sb2Te5 phase change compound with a machine-learned interatomic potential. npj Comput. Mater. 10, 33 (2024).
Google Scholar
Yang, M., Bonati, L., Polino, D. & Parrinello, M. Using metadynamics to build neural network potentials for reactive events: the case of urea decomposition in water. Catal. Today 387, 143–149 (2022).
Google Scholar
Guan, X., Heindel, J. P., Ko, T., Yang, C. & Head-Gordon, T. Using machine learning to go beyond potential energy surface benchmarking for chemical reactivity. Nat. Comput. Sci. 3, 965–974 (2023).
Google Scholar
Bonati, L. et al. The role of dynamics in heterogeneous catalysis: surface diffusivity and N2 decomposition on Fe (111). Proc. Natl. Acad. Sci. USA 120, e2313023120 (2023).
Google Scholar
David, R. et al. Arcann: automated enhanced sampling generation of training sets for chemically reactive machine learning interatomic potentials. Digit. Discov. 4, 54–72 (2025).
Google Scholar
Stocker, S., Gasteiger, J., Becker, F., Günnemann, S. & Margraf, J. T. How robust are modern graph neural network potentials in long and hot molecular dynamics simulations? Mach. Learn. Sci. Technol. 3, 045010 (2022).
Google Scholar
Batzner, S. et al. E (3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, 2453 (2022).
Google Scholar
Musaelian, A. et al. Learning local equivariant representations for large-scale atomistic dynamics. Nat. Commun. 14, 579 (2023).
Google Scholar
Batatia, I., Kovacs, D. P., Simm, G., Ortner, C. & Csányi, G. Mace: Higher order equivariant message passing neural networks for fast and accurate force fields. Adv. Neural Inf. Process. Syst. 35, 11423–11436 (2022).
Chanussot, L. et al. Open catalyst 2020 (OC20) dataset and community challenges. Acs Catal. 11, 6059–6072 (2021).
Google Scholar
Tran, R. et al. The open catalyst 2022 (OC22) dataset and challenges for oxide electrocatalysts. ACS Catal. 13, 3066–3084 (2023).
Google Scholar
Barroso-Luque, L. et al. Open materials 2024 (omat24) inorganic materials dataset and models. arXiv preprint arXiv:2410.12771 (2024).
Jain, A. et al. Commentary: The materials project: a materials genome approach to accelerating materials innovation. APL Mater 1, 011002 (2013).
Google Scholar
Deng, B. et al. Chgnet as a pretrained universal neural network potential for charge-informed atomistic modelling. Nat. Mach. Intell. 5, 1031–1041 (2023).
Google Scholar
Ghahremanpour, M. M. et al. The Alexandria Library, a quantum-chemical database of molecular properties for force field development. Sci. Data (2018).
Batatia, I. et al. A foundation model for atomistic materials chemistry. arXiv preprint arXiv:2401.00096 (2023).
Park, Y., Kim, J., Hwang, S. & Han, S. Scalable parallel algorithm for graph neural network interatomic potentials in molecular dynamics simulations. J. Chem. Theory Comput. 20, 4857–4868 (2024).
Google Scholar
Chen, C.-H., Chen, C., Ong, S. P. & Ong, S. P. A universal graph deep learning interatomic potential for the periodic table. Nat. Comput. Sci. 2, 718–728 (2022).
Google Scholar
Yang, H. et al. Mattersim: a deep learning atomistic model across elements, temperatures and pressures. arXiv preprint arXiv:2405.04967 arXiv: 2405.04967 (2024).
Póta, B., Ahlawat, P., Csányi, G. & Simoncelli, M. Thermal conductivity predictions with foundation atomistic models. arXiv.org arXiv: 2408.00755 (2024).
Peng, J., Damewood, J. K., Karaguesian, J., Lunger, J. R. & G’omez-Bombarelli, R. Learning ordering in crystalline materials with symmetry-aware graph neural networks. arXiv.org (2024).
Shiota, T., Ishihara, K., Do, T. M., Mori, T. & Mizukami, W. Taming multi-domain, -fidelity data: Towards foundation models for atomistic scale simulations. arXiv.org arXiv: 2412.13088 (2024).
Wang, Z. et al. Advances in high-pressure materials discovery enabled by machine learning. Matter Radiation Extremes 10, 033801 (2025).
Google Scholar
Kumar, A., Raghunathan, A., Jones, R., Ma, T. & Liang, P. Fine-tuning can distort pretrained features and underperform out-of-distribution. Int. Conf. Learn. Represent. (2022).
Kaur, H. et al. Data-efficient fine-tuning of foundational models for first-principles quality sublimation enthalpies. Faraday Discussions, 256, 120-138 (2025).
Radova, M., Stark, W. G., Allen, C. S., Maurer, R. J. & Bartók, A. P. Fine-tuning foundation models of materials interatomic potentials with frozen transfer learning. npj Comput Mater, 11, 237 (2025).
Deng, B. et al. Overcoming systematic softening in universal machine learning interatomic potentials by fine-tuning. arXiv.org arXiv: 2405.07105 (2024).
Zhuang, F. et al. A comprehensive survey on transfer learning. Proc. IEEE 109, 43–76 (2020).
Google Scholar
Hinton, G., Vinyals, O. & Dean, J. Distilling the knowledge in a neural network. NIPS Deep Learning and Representation Learning Workshop (2015).
Chen, M. S. et al. Data-efficient machine learning potentials from transfer learning of periodic correlated electronic structure methods: Liquid water at AFQMC, ccSD, and ccSD (t) accuracy. J. Chem. Theory Comput. 19, 4510–4519 (2023).
Google Scholar
Zaverkin, V., Holzmüller, D., Bonfirraro, L. & Kästner, J. Transfer learning for chemically accurate interatomic neural network potentials. Phys. Chem. Chem. Phys. 25, 5383–5396 (2023).
Google Scholar
Bocus, M., Vandenhaute, S. & Van Speybroeck, V. The operando nature of isobutene adsorbed in zeolite h-ssz-13 unraveled by machine learning potentials beyond DFT accuracy. Angew. Chem. Int. Ed. 64, e202413637 (2025).
Google Scholar
Röcken, S. & Zavadlav, J. Enhancing machine learning potentials through transfer learning across chemical elements. J. Chem. Inf. Model. 65, 7406–7414 (2025).
Google Scholar
Falk, J., Bonati, L., Novelli, P., Parrinello, M. & Pontil, M. Transfer learning for atomistic simulations using GNNs and kernel mean embeddings. Adv. Neural Inf. Process. Syst. 36, 29783–29797 (2023).
Muandet, K., Fukumizu, K., Sriperumbudur, B. & Schölkopf, B. Kernel mean embedding of distributions: a review and beyond. Found. Trends® Mach. Learn. 10, 1–141 (2017).
Google Scholar
Williams, C. & Seeger, M. Using the nyström method to speed up kernel machines. In Leen, T., Dietterich, T. & Tresp, V. (eds.) Advances in Neural Information Processing Systems (2000).
Rudi, A., Camoriano, R. & Rosasco, L. Less is more: Nyström computational regularization. In Cortes, C., Lawrence, N., Lee, D., Sugiyama, M. & Garnett, R. (eds.) Advances in Neural Information Processing Systems (NIPS, 2015).
Meanti, G., Carratino, L., Rosasco, L. & Rudi, A. Kernel methods through the roof: handling billions of points efficiently. In Advances in Neural Information Processing Systems 32 (NIPS, 2020).
Rahimi, A. & Recht, B. Random features for large-scale kernel machines. Advances in neural information processing systems 20 (2007).
Shi, L., Guo, X. & Zhou, D.-X. Hermite learning with gradient data. J. Comput. Appl. Math. 233, 3046–3059 (2010).
Google Scholar
Dhaliwal, G., Nair, P. B. & Singh, C. V. Machine learned interatomic potentials using random features. npj Computational Mater. 8, 7 (2022).
Google Scholar
Pham, N. & Pagh, R. Fast and scalable polynomial kernels via explicit feature maps. In Proceedings of the 19th ACM SIGKDD international conference on Knowledge discovery and data mining, 239–247 (Association for Computing Machinery, 2013).
Yu, F. X. X., Suresh, A. T., Choromanski, K. M., Holtmann-Rice, D. N. & Kumar, S. Orthogonal random features. Advances in Neural Information Processing Systems 29 (NIPS, 2016).
Rudi, A. & Rosasco, L. Generalization properties of learning with random features. Advances in Neural Information Processing Systems 30 (NeurIPS Proceedings, 2017).
Mei, S. & Montanari, A. The generalization error of random features regression: precise asymptotics and the double descent curve. Commun. Pure Appl. Math. 75, 667–766 (2022).
Google Scholar
Larsen, A. H. et al. The atomic simulation environment-a python library for working with atoms. J. Phys. Condens. Matter 29, 273002 (2017).
Google Scholar
Thompson, A. P. et al. LAMMPS—a flexible simulation tool for particle-based materials modeling at the atomic, meso, and continuum scales. Comp. Phys. Comm. 271, 108171 (2022).
Google Scholar
Jain, A. et al. Commentary: The materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Google Scholar
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Google Scholar
Owen, C. J. et al. Complexity of many-body interactions in transition metals via machine-learned force fields from the tm23 data set. npj Comput. Mater. 10, 92 (2024).
Google Scholar
Vandermause, J. et al. On-the-fly active learning of interpretable bayesian force fields for atomistic rare events. npj Comput. Mater. 6, 20 (2020).
Google Scholar
Lin, I.-C., Seitsonen, A. P., Tavernelli, I. & Rothlisberger, U. Structure and dynamics of liquid water from ab initio molecular dynamics: Comparison of blyp, pbe, and revpbe density functionals with and without van der waals corrections. J. Chem. theory Comput. 8, 3902–3910 (2012).
Google Scholar
Montero de Hijes, P., Dellago, C., Jinnouchi, R., Schmiedmayer, B. & Kresse, G. Comparing machine learning potentials for water: Kernel-based regression and behler-parrinello neural networks. J. Chem. Phys. 160, 114107 (2024).
Google Scholar
Kovács, D. P. et al. MACE-OFF: Short-Range Transferable Machine Learning Force Fields for Organic Molecules. J. Am. Chem. Soci. 147, 17598–17611 (2025).
Google Scholar
Eastman, P. et al. Spice, a dataset of drug-like molecules and peptides for training machine learning potentials. Sci. Data 10, 11 (2023).
Google Scholar
Hodgson, A. & Haq, S. Water adsorption and the wetting of metal surfaces. Surf. Sci. Rep. 64, 381–451 (2009).
Google Scholar
Nagata, Y., Ohto, T., Backus, E. H. & Bonn, M. Molecular modeling of water interfaces: From molecular spectroscopy to thermodynamics. J. Phys. Chem. B 120, 3785–3796 (2016).
Google Scholar
Mikkelsen, A. E., Schiotz, J., Vegge, T. & Jacobsen, K. W. Is the water/pt (111) interface ordered at room temperature? J. Chem. Phys. 155, 224701 (2021).
Perego, S. & Bonati, L. Data efficient machine learning potentials for modeling catalytic reactivity via active learning and enhanced sampling. npj Comput. Mater. 10, 291 (2024).
Google Scholar
Hospedales, T., Antoniou, A., Micaelli, P. & Storkey, A. Meta-learning in neural networks: a survey. IEEE Trans. Pattern Anal. Mach. Intelligence 44, 5149–5169 (2021).
Falk, J. I. T., Ciliberto, C. & Pontil, M. Implicit kernel meta-learning using kernel integral forms. In Uncertainty in Artificial Intelligence, 652–662 (PMLR, 2022).
Kelvinius, F. E., Georgiev, D., Toshev, A. P. & Gasteiger, J. Accelerating molecular graph neural networks via knowledge distillation. Adv. Neural Inf. Process. Sys. 36, 257611–25792 (2023).
Matin, S. et al. Teacher-student training improves accuracy and efficiency of machine learning interatomic potentials. Digital Discovery, Advance Article (2025).
Gardner, J. L. A. et al. Distillation of atomistic foundation models across architectures and chemical domains. arXiv.org arXiv: 2506.10956 (2025).
Amin, I., Raja, S. & Krishnapriyan, A. Towards fast, specialized machine learning force fields: Distilling foundation models via energy hessians. The Thirteenth International Conference on Learning Representations (2025).
Sipka, M., Erlebach, A. & Grajciar, L. Constructing collective variables using invariant learned representations. J. Chem. Theory Comput. 19, 887–901 (2023).
Google Scholar
Pengmei, Z., Lorpaiboon, C., Guo, S. C., Weare, J. & Dinner, A. R. Using pretrained graph neural networks with token mixers as geometric featurizers for conformational dynamics. J. Chem. Phys. 162, 044107 (2025).
Google Scholar
Trizio, E., Rizzi, A., Piaggi, P. M., Invernizzi, M. & Bonati, L. Advanced simulations with plumed: Opes and machine learning collective variables. arXiv preprint arXiv:2410.18019 (2024).
Rudin, W.Fourier analysis on groups (Courier Dover Publications, 2017).
Kresse, G. & Hafner, J. Ab initio molecular dynamics for liquid metals. Phys. Rev. B 47, 558 (1993).
Google Scholar
Kresse, G. & Hafner, J. Ab initio molecular-dynamics simulation of the liquid-metal–amorphous-semiconductor transition in germanium. Phys. Rev. B 49, 14251 (1994).
Google Scholar
Kresse, G. & Joubert, D. From ultrasoft pseudopotentials to the projector augmented-wave method. Phys. Rev. b 59, 1758 (1999).
Google Scholar
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169 (1996).
Google Scholar
Kresse, G. & Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Computational Mater. Sci. 6, 15–50 (1996).
Google Scholar
Kresse, G. & Hafner, J. Norm-conserving and ultrasoft pseudopotentials for first-row and transition elements. J. Phys.: Condens. Matter 6, 8245 (1994).
Google Scholar
Paszke, A. Pytorch: An imperative style, high-performance deep learning library. Adv. neural inf. process. sys. 32, (2019).
link