Accurate RNA 3D structure prediction using a language model-based deep learning approach

Automated end-to-end platform for RNA 3D structure prediction
The development of RhoFold+ was guided by RNA-specific knowledge and the limitations of existing RNA 3D structure data. To build our training dataset, we curated all available RNA 3D structures from the PDB, using the BGSU representative sets of RNA structures (version 2022-04-13)24. We focused on single-chain RNAs and reduced redundancy by clustering sequences with Cd-hit25 at an 80% sequence similarity threshold, resulting in 782 unique sequence clusters from 5,583 RNA chains. These RNA sequences were then processed through our pipeline, RhoFold+. First, the sequences were transformed using RNA-FM, our large RNA language model, to extract evolutionarily and structurally informed embeddings. Concurrently, MSAs were generated by searching through extensive sequence databases. The embeddings and MSA features were then fed into our transformer network, Rhoformer, and iteratively refined for ten cycles. Following this, our structure module employed a geometry-aware attention mechanism and an invariant point attention (IPA) module to optimize local frame coordinates and torsion angles for key atoms in the RNA backbone. Structural constraints, such as secondary structure and base pairing, were applied after reconstructing the full-atom coordinates (Fig. 1a and detailed discussion in Supplementary information). After developing RhoFold+, we rigorously benchmarked and evaluated its performance across a broad range of tests (Fig. 1b).
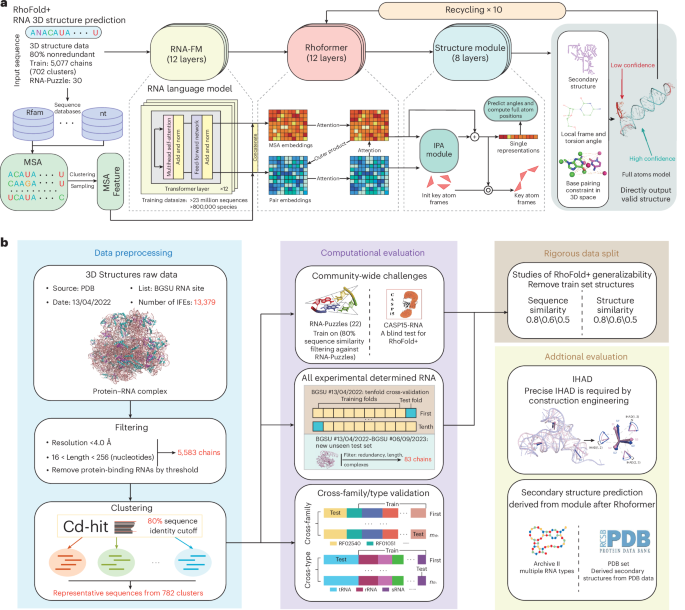
a, The architecture of RhoFold+, a fully automated and differentiable end-to-end approach to de novo RNA 3D structure prediction from the sequence. Using an RNA language model (RNA-FM) pretrained on 23,735,169 unannotated RNA sequences and several deep learning modules—including an IPA module that models 3D positions—RhoFold+ can generate valid and largely accurate RNA 3D structures of interest typically within ~0.14 s (without MSA searching). init, initialized; norm, normalize. b, The preprocessing step of RhoFold+ to extract all available nonredundant single-stranded RNA 3D structures from the PDB database. IFE, integrated functional element. RhoFold+ is comprehensively benchmarked on community-wide challenges including RNA-Puzzles targets and CASP15 natural RNA targets, and on all available experimentally determined RNA 3D structures. RhoFold+ also demonstrates high accuracy in cross-validation experiments, as well as generalizability to unseen, newly determined RNA structures and unseen RNA families and types in cross-family and cross-type validation experiments. Data split evaluations reveal that RhoFold+ does not overfit its training set. RhoFold+ is also capable of predicting secondary structures and parameters that are useful for construct engineering.
Benchmarking RhoFold+ on RNA-Puzzles
We performed a comprehensive retrospective comparison between RhoFold+ and other existing computational methods on two previously held community-wide challenges: RNA-Puzzles and CASP15. We first used the results from the RNA-Puzzles26,27,28,29,30 competition, where the submissions were produced and optimized by human knowledge or computational methods. Importantly, here RhoFold+ was trained using nonoverlapping training data with respect to the RNA-Puzzles targets tested (Methods). We conducted preprocessing to obtain 24 single-chain RNA targets and excluded RNA complexes. This set of RNA targets contained two puzzles (PZs), PZ34 and PZ38, that were introduced after our development of RhoFold+ (Fig. 2a and Supplementary Fig. 3) and thus served as a blind test. After collecting the predictions of other methods from the official server ( we found that the performance of RhoFold+ surpassed that of all other methods, including FARFAR2/ARES, on nearly all targets, except for PZ24. Notably, RhoFold+ outperformed the second-best method on more than half of the targets by ~4 Å r.m.s.d. On 17 targets, RhoFold+ achieved r.m.s.d. values of <5 Å, and only one target exhibited an r.m.s.d. of >10 Å (Fig. 2a and Supplementary Table 5). As a whole, RhoFold+ produced an average r.m.s.d. of 4.02 Å, 2.30 Å better than that of the second-best model (FARFAR2: top 1%, 6.32 Å). Assessed using the template modeling (TM) score31, RhoFold+ achieved an average of 0.57 (Supplementary Table 5), higher than the scores of other top performers (0.41 and 0.44).
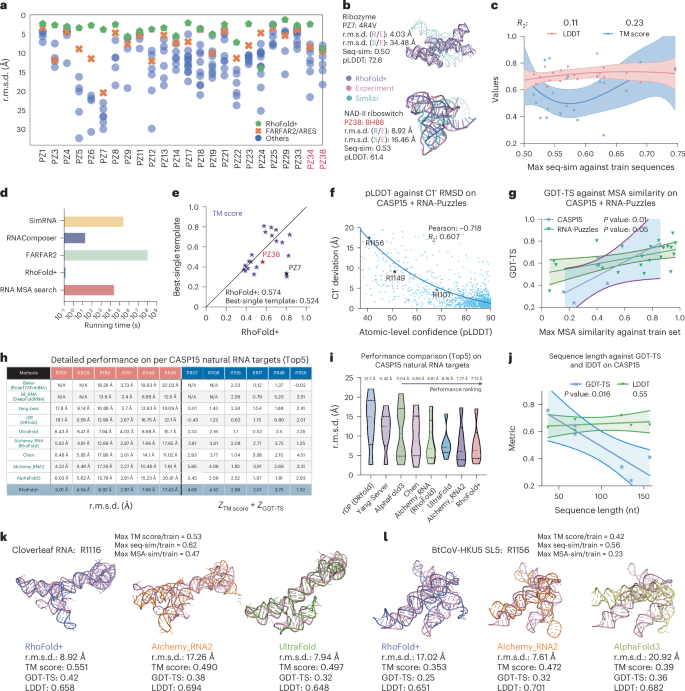
a, The r.m.s.d. performance scatter plot of RhoFold+ and other methods across 24 nonoverlapping, nonredundant RNA-Puzzles targets. Each point represents a predicted model from the specific method. b, Visualization of RNA-Puzzles 7 and 38. In addition to the aligned RhoFold+ prediction, we show the most similar training structure with respect to each target, suggesting that RhoFold+ neither overfits the training set nor simply reproduces the most similar structure to the target. Seq-sim, sequence similarity. c, Regression plot of the TM score and LDDT of RhoFold+ predictions against the maximum sequence similarity among all the training sequences, across all RNA-Puzzles targets. Each point represents an RNA-Puzzles target. d, The running time comparison for different methods. e, Comparison of RhoFold+ predictions against the respective best single templates from our training set across all RNA-Puzzles targets. f, A regression plot for the r.m.s.d. of against atom-level pLDDT across all RNA-Puzzles and CASP15 targets. g, A regression plot for structure GDT-TS against MSA similarity across all RNA-Puzzles and CASP15 targets. h, A detailed performance comparison for CASP15 natural RNA targets. The pink columns record detailed r.m.s.d. values and the blue columns record the sum of Z-scores for the GDT-TS and TM score. Entries missing officially reported CASP15 data are marked as N/A; Yang-Sever and Chen are CASP15 registered groups. i, A comparison of RhoFold+’s average performance against the average reported performance of CASP15 groups and published works on CASP15 natural RNA targets. j, A regression plot for the structure GDT-TS and LDDT against sequence length across all CASP15 targets. The central curve in c, g and j represents the fit regression model, while the two surrounding curves indicate the 95% percentile intervals. k, A comparison of RhoFold+ predictions against AIchemy_RNA2 and UltraFold on the R1116 target from CASP15. MSA-sim, MSA profile similarity. l, For the R1156 target, showing a RhoFold+ potential failure case, involving incorrect stacking patterns and orientations.
Source data.
To show that the promising results on RNA-Puzzles did not arise from overfitting, we studied whether the sequence similarity between the test set and our training data was substantially positively correlated with the performance of RhoFold+, as measured by the TM score and the local distance difference test (LDDT), a superposition-free score that evaluates local distance differences for all atoms in a model32,33. Such a correlation was previously found in protein structure prediction13, yet here we found that R2 values, which represent whether the slope is significantly nonzero, were 0.23 for the TM score and 0.11 for the LDDT (Fig. 2b,c), indicating no significant correlation between model performance and the similarity of our training and testing sets. These results suggest that RhoFold+ can generalize in predicting accurate RNA structures. A case study of a representative RNA-Puzzles target, PZ7 (a 186-nucleotide-long Varkud satellite ribozyme RNA), exemplifies this finding. Here, the structure of the most similar RNA in the training set differed substantially from the structure of PZ7 (Fig. 2b): the r.m.s.d. between these structures was 34.48 Å. As another example, PZ38 exhibited the highest sequence similarity of 53% with respect to all RNAs in our training set, and the r.m.s.d. between the structure of the most sequence-similar RNA and PZ38 was 16.46 Å (Fig. 2b). This was larger than the r.m.s.d. of 8.92 Å between PZ38 and the RhoFold+ prediction.
To test the ability of RhoFold+ to generalize for structure-dissimilar (in addition to mainly sequence-dissimilar) targets, we sought to determine whether the predictions of RhoFold+ could surpass the best single template (the most structurally similar model) in the training set for a given query. To investigate this, we compared the TM scores between our predictions and experimentally determined structures against the TM scores between the best single templates and experimentally determined structures across all RNA-Puzzles. For the majority of puzzles, RhoFold+ produced predictions with a higher global similarity and an average TM score of 0.574, surpassing the best single template by 0.05 (Fig. 2e and Supplementary Table 13). It is important to highlight that for proteins, surpassing the best single template required substantial progress. Indeed, it was only during CASP14 that computational methods outperformed the best single template. Although RhoFold+ generated considerably more accurate predictions than other methods under the conventional sequence similarity data splitting paradigm, we further tested the adaptability of RhoFold+ by eliminating 3D structures from the training set whose TM score, with respect to any target, surpassed a specified threshold (Supplementary Fig. 6 and Supplementary Tables 6 and 10). Even under this more demanding condition, RhoFold+ continued to exhibit a promising performance (Supplementary Table 10).
In applying computational models to large-scale, real-world settings, speed is often a top priority. In addition to generating largely accurate folding results, we found that RhoFold+ is fast, with typical RNA-Puzzles predictions completed within ~0.14 s (Fig. 2d). In contrast, other approaches, including SimRNA12, FARFAR210 and RNAComposer34, exhibited significantly longer running times, probably due to the large-scale sampling processes employed by these methods (Fig. 2d).
Benchmarking RhoFold+ on CASP15 targets
As RNA-Puzzles was first released over a decade ago26, we next used RhoFold+ to predict RNA targets from the more recent CASP15 (refs. 35,36). We focused on CASP15’s six natural RNA targets (Fig. 2h and Supplementary Fig. 4). Artificially designed targets, which fell outside the expected domain of application for RhoFold+, were not included: in particular, the excluded targets were characterized by their lack of homology and divergence from our training set or their being RNA–protein complexes. We followed the CASP15 guidelines, which specified that participating teams were permitted to submit up to five models. Utilizing different, randomly sampled MSAs (Methods), we modeled five candidate structures for each target using RhoFold+ and considered only the highest-performing prediction (Supplementary Table 6).
Several top-ranking CASP15 groups and recent published works on RNA 3D structure prediction17,18,20,21,23 were included in our benchmarking. Particularly, CASP15 groups were divided into two categories, ‘server’ and ‘expert’, depending on whether or not human expert knowledge and fine tuning were used. Regardless of the category, many CASP15 groups employed computational pipelines that were based on comparative or statistical learning for natural targets, thus allowing us to assess the learning capability of RhoFold+. Our preliminary model, AIchemy_RNA (RhoFold), was a participant in the ‘expert’ category. Building on RhoFold, RhoFold+ represents a fully automated and end-to-end pipeline that is more similar to participants in the ‘server’ category. Here, we found that RhoFold+ outperformed RhoFold on CASP15’s natural RNA targets by an average r.m.s.d. of ~1 Å. Furthermore, RhoFold+ outperformed other methods whose predictions were available for all six natural RNA targets, including the first-ranked AIchemy_RNA2, the second-ranked Chen method and other computational methods, including DRfold23, DeepFoldRNA17, AlphaFold321 and trRosettaRNA18 (Fig. 2h,i). Although RhoFold+ outperformed AIchemy_RNA2 marginally by 0.06 Å (average r.m.s.d.; Fig. 2i), AIchemy_RNA2 required expert knowledge. Additionally, RhoFold+ demonstrated accuracy comparable to each top-performing method on almost every natural RNA target, with the exception of R1156 (Fig. 2h).
Following CASP15’s assessment approach36, we also computed Z-scores for the predictions from all participating groups. CASP15 prioritized the TM score and the global distance test-total score (GDT-TS), which evaluates both overall structure similarity and local alignment, leading us to assess these models based on the cumulative Z-scores of these metrics (Fig. 2h). On the six natural RNA targets and among the subset of all CASP15 participants ranked on these specific targets, RhoFold (AIchemy_RNA) was fourth, while the performance of RhoFold+ was on par with that of AIchemy_RNA2 (with a difference of 0.4 in the Z-score) and surpassed that of other methods. In a detailed analysis of performance on specific targets, we found that, for target R1108, RhoFold+ achieved the best Z-score and r.m.s.d. Interestingly, RhoFold+ also attained the best Z-score for R1116, although the r.m.s.d. was ~1 Å higher than that of UltraFold (other methods produced predictions with significantly lower accuracy, all with r.m.s.d. >10 Å). Upon further investigation, we found that, while UltraFold outperformed RhoFold+ on this metric by producing accurate local predictions, the predicted global structure was less accurate, as evidenced by a TM score of 0.497 and a GDT-TS score of <0.4. In contrast, RhoFold+ inaccurately predicted a helix angle, resulting in an r.m.s.d. of 8.92 Å, but its correctly predicted topology resulted in a higher TM score of >0.55. For this target, AIchemy_RNA2 incorrectly predicted the stem stackings and RNA topology, resulting in a high r.m.s.d. of 17.26 Å and a TM score of ~0.49. Notably, the RhoFold+ prediction for R1116 did not arise from overfitting, as indicated by the low maximum structural similarity (TM score) and maximum sequence similarity of R1116 with respect to the training set (Fig. 2k and Supplementary Table 6).
We also looked into targets where RhoFold+ may achieve reduced performance and found that higher MSA quality correlated with better performance. While RhoFold+ accurately predicted local structural topologies, it struggled with aligning helices, particularly at junctions. This discrepancy may be due to the dynamic and flexible nature of RNA junctions, which often adopt multiple conformations37,38,39, making them challenging for fully automated models to represent accurately (Fig. 2k,l and detailed discussion in Supplementary information).
Factors influencing prediction accuracy
Building on the findings above, we performed a more comprehensive study involving all CASP15 natural RNAs and RNA-Puzzles targets. We observed that the prediction accuracy of RhoFold+ is sensitive toward the query’s MSA profile similarity (Supplementary information) against the training set (Fig. 2g) and the complexity of RNA structures (query length; Fig. 2j). Additionally, predicted LDDT (pLDDT) scores were found to correlate with the confidence of RhoFold+, providing a useful metric for identifying regions with lower prediction accuracy, especially in more complex or less homologous queries (Fig. 2f and detailed discussion and analysis in Supplementary information).
Benchmarking RhoFold+ on all determined RNA 3D structures
After benchmarking RhoFold+ with RNA-Puzzles and CASP15, we next evaluated RhoFold+ in greater detail using all experimentally determined RNA structures, as defined by the BGSU representative sets of RNA structures (preprocessed to remove redundancy). To further study the performance of RhoFold+, we performed tenfold cross-validation by iteratively masking 80 sequence clusters for validation and leaving 702 sequence clusters for training. We found that the performance of RhoFold+ across all RNA structures was robust regardless of the train–test data split and fairly consistent across all folds (Fig. 3a–c). Slight variations in TM score might be caused by challenging targets such pseudoknot cases in Fold2 and Fold7 similar to PZ24 (Fig. 3c,e), and we expect that the predictions of RhoFold+ on such targets could be improved if secondary structure constraints were provided. Also, during our cross-validation test, the accurate predictions of RhoFold+ were not due to merely mimicking the most sequence-similar training data (Fig. 3b,d,e). A plot of the r.m.s.d. against the sequence length shows that r.m.s.d. values were largely distributed below 10 Å, independent of the sequence length (Fig. 3a). Outliers with r.m.s.d. >20 Å were more likely to occur for sequences longer than 200 nt, where we expect further improvement by more tuning on long RNAs (detailed discussion in Supplementary information).
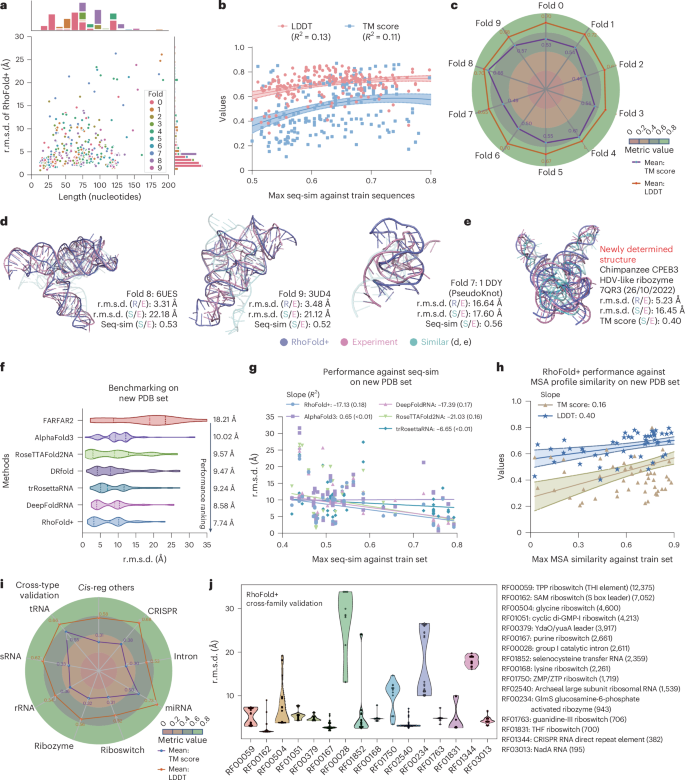
a, A plot of r.m.s.d. values against sequence length for all cross-validation experiments. Each point represents an RNA structure and is colored according to the cross-validation fold. b, A regression analysis for each prediction’s TM score (blue) and LDDT (pink) against the maximum sequence similarity with respect to all training data. Each point represents an RNA structure. c, The average TM score and LDDT for each fold. d, Visualization of two representative riboswitch structures, 6UES and 3UD4, and a pseudoknot 1DDY (pink), along with the corresponding RhoFold+ predictions (slate) and the training RNA structures with the highest sequence similarity (cyan). In a–d, the tenfold cross-validation of RhoFold+ using all experimentally determined RNA structures is shown. e, Visualization of a newly determined RNA structure, 7QR3, an hepatitis delta virus (HDV)-like ribozyme, which has a low structural similarity with respect to the training set, but whose structure (pink) is accurately predicted by RhoFold+ (slate). The most similar structure, 7DLZ, is shown in cyan. f, A comparison of average r.m.s.d. values generated by RhoFold+ and other methods on the new PDB set, a set of 76 newly determined solo RNA structures. g, A regression plot of the prediction r.m.s.d. values against maximum sequence similarity to the training set for RhoFold+ and other baseline methods. h, A regression plot of the correlation between the RhoFold+ predictions TM score/LDDT and the maximum MSA profile similarity against the training set. The central curve in b and h represents the fit regression model, while the two surrounding curves indicate the 95% percentile intervals. i, An overview of cross-type validation performance of RhoFold+ measured by LDDT and TM score. All structures in the type used for validation were masked during model training. sRNA, small RNA. j, A violin plot of RhoFold+ r.m.s.d. values in the cross-family validation. Here, all the structures in a family to be tested were masked during model training and RhoFold+ accurately predicted RNA structures from most unseen families. The numbers of sequences in each family are shown in parentheses.
Source data.
As a further evaluation of the capabilities of RhoFold+, we considered the model’s performance on newly determined RNA single-stranded structures released subsequent to the compilation of our training dataset. This approach acted as an additional blind test, similar to the CASP15 competition. We included comparisons against FARFAR2 and recent deep learning methods17,18,21,23, all of which have inference code and/or servers available and some of which also participated in CASP15 (Methods). RhoFold+ outperformed all benchmarked models, achieving the highest average accuracy as measured by r.m.s.d. RhoFold+ produced an average r.m.s.d. of 7.74 Å, which was approximately 0.8 Å and 10.5 Å better than the second-ranked DeepRNAFold and the lowest-ranked FARFAR2, respectively. Notably, on average, RhoFold+ also outperformed AlphaFold3 and RoseTTAFold2NA by approximately 2.2 Å and 1.8 Å, respectively (Fig. 3f and detailed discussion in Supplementary information). These results were consistent with the performance observed in our previous benchmark on CASP15, suggesting that RhoFold+ accurately generalizes to newly determined structures not seen in our training set. Furthermore, these results support that AlphaFold3 and RoseTTAFold2NA, which are designed to predict biomolecular complexes, do not perform as well as RhoFold+ when applied to single RNA molecules. Further examining sequence and structural similarities to our training set reveals that RhoFold+ maintained strong performance even with sequence similarities below 0.5 (Fig. 3g), and the TM score was greatly influenced by MSA profile similarity while local accuracy (LDDT) remained high and robust (Fig. 3h). Additionally, RhoFold+ demonstrated strong generalizability, accurately folding structures such as 7QR3 despite its low similarity to the closest training template, 7DLZ (TM score of 0.40, r.m.s.d. of 16.45 Å; Fig. 3e).
RhoFold+ generalizes to unseen RNA types and families
Having demonstrated that RhoFold+ can generalize to predicting RNA structures with divergent sequence similarities, structural similarities and dates of release, we next investigated the ability of RhoFold+ to handle different RNA types and families defined by expert knowledge. In particular, RNA types and families—such as those curated in Rfam40—are often classified manually based on factors including function, structure and co-evolutionary information. Addressing the challenge of generalizing to different RNA types and families may be considerably more demanding for deep learning methods such as RhoFold+ as such a task requires larger domain shifts.
We benchmarked the cross-type performance of RhoFold+ by training the model on a subset of all RNA types while testing on the others. RhoFold+ showed robustness across RNA types. Though struggling with introns and riboswitches, it performed well on transfer RNA (tRNA) and micro RNA (miRNA) types, achieving TM scores up to 0.73 (Fig. 3i). When compared with FARFAR2, RhoFold+ outperformed it across all RNA types, particularly in tRNAs and ribosomal RNAs (rRNAs), with smaller margins for riboswitches (detailed discussion in Supplementary information). For cross-family tests, RhoFold+ achieved an average r.m.s.d. of 6.69 Å (Fig. 3j), but struggled with complex families such as group I introns (RF00028). This difficulty is consistent with challenges observed in cross-type tests, such as for complex RNA types such as introns and CRISPR RNA elements (RF01344). These elements interact with various proteins and enzymes, and focusing solely on RNA structure without considering these interactions may limit the prediction accuracy (detailed discussion in Supplementary information). Overall, these tests demonstrate the ability of RhoFold+ to generalize across unseen RNA types and families, though challenges remain for complex structures and datasets with limited available data.
RhoFold+ predicts secondary structures and substructures
RhoFold+ can accurately predict RNA 3D structures, but the limited number of experimentally determined RNA structures and types makes it difficult to understand the space of all possible RNA folds. This is particularly true for complicated and large RNA types, including internal ribosomal entry sites, introns, synthetic RNAs and long noncoding RNAs. RNA secondary structures, however, can be more easily determined in experiments and accurate secondary structure predictions can supplement the predictions of 3D structures, offering valuable insights into RNA folding and function. Therefore, we adapted RhoFold+ to predict secondary structures as well. As RhoFold+ was designed to predict RNA 3D structures, we incorporated a postprocessing module that utilizes the features retrieved from RhoFold+’s Rhoformer to predict secondary structures (since Rhoformer’s features show attention maps highly aligned with the contact maps; Supplementary Fig. 8 and Supplementary Table 14). This module takes into account the same structural information as the module performing 3D reconstruction but operates under distinct geometric and biological constraints imposed to predict secondary structure.
We benchmarked the performance of RhoFold+ on newly determined PDB structures (the ‘new PDB set’) and the ArchiveII dataset41, which includes secondary structure information for diverse RNAs. On the new PDB set, RhoFold+ outperformed UFold41 by 0.035 in the average F1 score (Fig. 4a), even when UFold was trained on all available data (PDB and bpRNA-1M, a database with over 100,000 annotated RNA secondary structures). On the ArchiveII dataset comprising 2,975 RNA samples, RhoFold+ also outperformed other secondary structure prediction methods (Fig. 4b), particularly on larger RNA types (Fig. 4c). For instance, it achieved an F1 score of 0.60 on structured domains in the dengue virus transcriptome (Supplementary Table 19), aligning with results from mutational profiling (RING-MaP)42,43. Similarly, the strong performance of RhoFold+ did not stem from mimicking training data, as it maintained an F1 score of ~0.7 even when sequence similarity dropped below 50% (Fig. 4e), and achieved a perfect F1 score of 1.0 on the CASP15 target R1117 (Fig. 4f). These results suggest that RhoFold+ not only excels in predicting 3D structures, but also generates rich, meaningful representations that enable state-of-the-art secondary structure prediction.
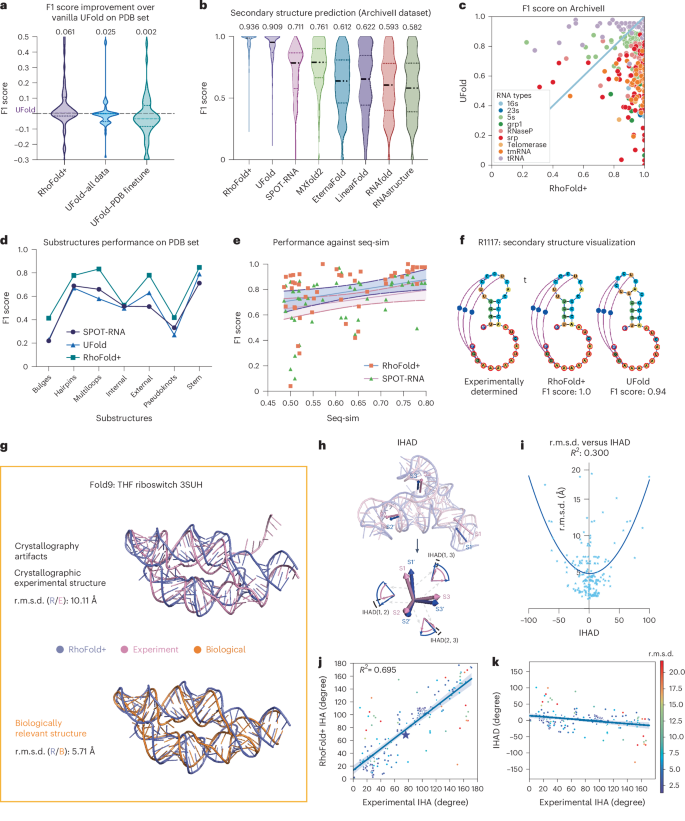
a, F1 score comparison against multiple configurations of UFold on the PDB set. Here, a version of UFold trained on bpRNA is also presented as a baseline, to evaluate the improvement in terms of F1 score. b, The F1 score distribution of various methods on the ArchiveII dataset. Average scores are indicated at the top of the plot. c, F1 score comparison between RhoFold+ and UFold on the ArchiveII dataset. Each point represents an RNA structure and is colored according to its RNA type. srp, signal recognition particle RNA; tmRNA, transfer-messenger RNA. d, F1 score comparison of RhoFold+ versus UFold and SPOT-RNA on RNA substructures in the new PDB set. e, F1 score comparison of RhoFold+ versus UFold and SPOT-RNA against sequence similarity of RNA structures in the new PDB set. f, Visualization of a CASP15 RNA target where RhoFold+ predicted the correct secondary structures including pseudoknots. g, Visualization of a swapped dimer, tetrahydrofolate (THF) ribozyme, 3SUH, for which the RhoFold+ prediction (purple) resembles the biologically meaningful structure (orange) instead of the crystallographic artifact found in the PDB (pink). h, Visualization showing the definition of the IHAD, which is the difference between the IHAs derived from the RhoFold+ prediction and the experimentally determined structure. i, Regression analysis between the IHAD and r.m.s.d. of the RhoFold+ predictions. Each point represents an RNA. j, Comparison between the IHAs derived from the RhoFold+ predictions against those from experimental structures. Each point represents an angle instance and is colored according to the r.m.s.d. between the experimental structure containing the angle and the structure predicted by RhoFold+. k, A plot of the IHAD against experimentally determined IHA values. The coloring is the same as in j. The central curve in e, j and k represents the fit regression model, while the two surrounding curves indicate the 95% percentile intervals.
Source data.
We further evaluated substructures within RNA secondary structures, finding that RhoFold+ consistently outperformed SPOT-RNA44 and UFold41 across all substructures, with the most significant improvements in multiloops and external loops, while internal loops and pseudoknots showed similar performance across methods (Fig. 4d). These results underscore the potential capability of RhoFold+ in predicting RNA secondary structures and enhancing our understanding of RNA function.
Correcting artifacts and IHA prediction
As RhoFold+ accurately predicts RNA structures at both the secondary and tertiary levels, we asked whether we could leverage RhoFold+ for experimental efforts. Toward this, we investigated two use cases of RhoFold+: (1) for correcting experimental structural artifacts and (2) for guiding RNA construct engineering.
X-ray crystallography is widely used to resolve RNA 3D structures, but it can introduce artifacts such as domain-swapped dimers45, potentially misleading machine learning models that do not generalize well. In one case, the RhoFold+ prediction for 3SUH initially yielded a high r.m.s.d. of 10.11 Å compared with the PDB structure. However, further analysis revealed that the crystal structure involved a domain-swapped dimer. When comparing the RhoFold+ prediction with the inferred monomeric structure, the r.m.s.d. improved to 5.71 Å, indicating RhoFold+ accurately predicted the biologically relevant structure (Fig. 4g). Similar findings were also observed for the ZTP riboswitch46 (Supplementary Fig. 9), suggesting that RhoFold+ can effectively correct for such experimental artifacts.
When comparing experimental data with RNA 3D models, additional geometric metrics, such as interhelical angles (IHAs), can provide insights beyond standard global alignment measures such as r.m.s.d., LDDT and TM score. IHAs, which can be estimated using experimental methods, are useful for validating predicted models and guiding RNA nanostructure design. We introduced the IHA difference (IHAD) as a metric to benchmark the predictions of RhoFold+ (Fig. 4h and Supplementary information), finding that IHAD can reveal discrepancies in stem orientations that are not captured by r.m.s.d. alone (Fig. 4i). Our analysis shows that RhoFold+ generally predicted stem directions accurately (Fig. 4j,k), though performance decreased for IHAs near 0° or 180°, probably due to underfitting of parallel stems in large and complex structures (Fig. 4k and detailed discussion in Supplementary information). We further demonstrated the practical application of IHAs by predicting values for RNA constructs such as the FMN riboswitch and the P4–P6 domain from the Tetrahymena group I intron (Supplementary Fig. 9).
Ablation studies and generation of multiple predictions
Given the high accuracy and speed of RhoFold+, we finally conducted ablation studies to understand which components and information are important to the RhoFold+ predictions. The architectural components we investigated included four different modules (Fig. 5a and Methods). Ablation studies were performed on 138 PDB targets (collected between April 2022 and December 2023) with sequence similarities below 80% to our training set and lengths ranging from 16 to 300 nt (the ‘Ablation set’). By removing each RhoFold+ component, we observed that all contributed to improving the performance, with the MSA module being the most critical, followed by the RNA-FM language model (Fig. 5a). The RNA-modified version of AlphaFold2, without the MSA module, performed worse than RhoFold+ (Fig. 5a). Notably, removing RNA-FM led to a sharper performance decline for dissimilar sequences (Fig. 5b), and the RNA-FM module seemed to compensate for the loss of the MSA module, maintaining higher TM scores (Fig. 5c). Additionally, removing the recycling module most significantly affected predictions for longer sequences, probably due to its role in effectively deepening the model (Supplementary Fig. 7 and detailed discussion in Supplementary information).
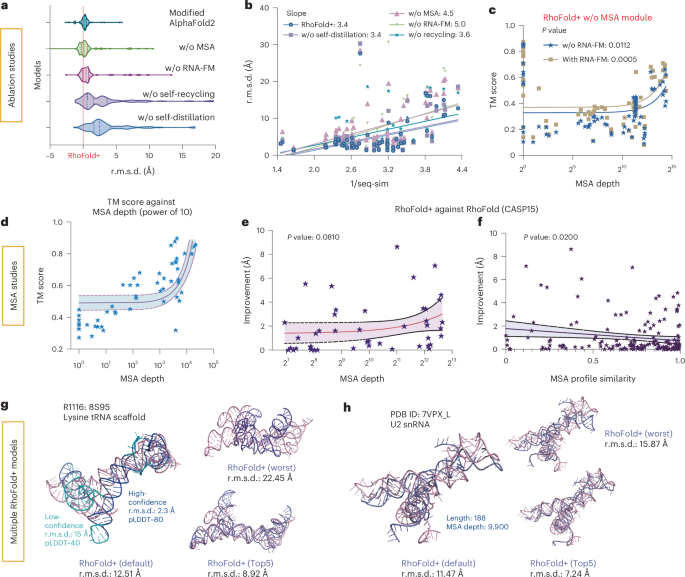
a, Ablation studies of RhoFold+ without (w/o) corresponding modules in RhoFold+ with performance measured by r.m.s.d. b, A regression analysis for prediction accuracy (measured by r.m.s.d.) against the reciprocal of sequence similarity. c, A regression analysis of the TM score against MSA depth for the ablation study of the RNA-FM module. Note that the x axis is log scaled. d, A plot of prediction accuracy (measured by the TM score) against MSA depth. e, A plot of the improvement of RhoFold+ against RhoFold (measured by r.m.s.d.) across different MSA depths. f, A plot of the improvement of RhoFold+ against RhoFold (measured by r.m.s.d.) across different MSA profile similarities. The central curve in e and f represents the fit regression model, while the two surrounding curves indicate the 95% percentile intervals. g, Visualization of a CASP15 target where RhoFold+ produces an r.m.s.d. of 12.51 Å, but improves by 8.92 Å using the Top5 prediction from MSA sampling. h, Visualization of a newly determined RNA structure where the r.m.s.d. of RhoFold+ improves by 7.92 Å using Top5 prediction from MSA sampling.
Source data
These findings are consistent with our results for CASP15’s natural RNA targets and RNA-Puzzles, where MSA quality significantly impacts predictions. We also explored how the number of sequences in the extracted MSA influences accuracy. While RhoFold+ is limited to 256 MSAs due to training constraints, this limit did not compromise its effectiveness. A key enhancement in RhoFold+ is its ability to generate multiple predictions by sampling or clustering from a fixed number of MSAs, allowing for broader prediction selection and improved outcomes. Performance on RNA-Puzzles showed an inverse correlation with reduced MSA counts, with a marked improvement when the MSA number exceeded 100 (Fig. 5d), indicating that a larger MSA pool enhances model optimization (detailed discussion in Supplementary information). With this expanded MSA sampling, the lowest r.m.s.d. of the RhoFold+ Top5 predictions significantly decreased compared with RhoFold, correlating positively with increased MSA depth and yielding an up to 10 Å improvement (Fig. 5e). This improvement was more pronounced when the MSA profile similarity between the query and training sequences was high, resulting in smaller gains when similarity was already strong (Fig. 5f). Overall, additional MSA sampling is crucial for high performance, as demonstrated for CASP15 target R1116 and PDB 7VPX_L (Fig. 5g,h).
link