AI-guided competitive docking for virtual screening and compound efficacy prediction
Overview
The first part of the results evaluated how diffusion-based co-folding models differentiate true inhibitors from inactive compounds, using a dataset of 16 protein targets in addition to the more complex multi-site DNA gyrase. The second part explores the use of diffusion-based tools for ranking inhibitors, introducing a competitive docking strategy applied to all targets. Finally, we present two applications of this competitive docking approach to DNA gyrase: an All-at-Once virtual screening method for hit identification and a strategy for designing de novo inhibitors with improved predicted potency.
Pose convergence can help identify real inhibitors
We used 16 protein targets with diverse biological functions as benchmarks to assess whether denoising diffusion-based models can distinguish true binders from false positives. After reviewing the reference crystal structures for these benchmark proteins (Table S1), we predicted binding poses for several inhibitors lacking experimental structures. Additionally, we included 28 unrelated “off-target” compounds for each benchmark (Table S2). For a given protein target, the off-target set consisted of one inhibitor from each of the other targets in the study, along with compounds that typically bind to proteins entirely unrelated to those analyzed here.
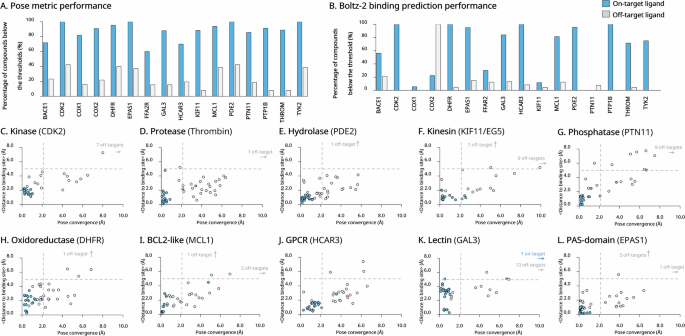
We assessed docking specificity using two criteria: (i) how closely ligands remained within the binding site across predicted models, and (ii) how consistent their poses were with each other (pose convergence), measured by the average RMSD. Overall, true inhibitors bound within approximately 5 Å of the binding pocket and exhibited strong convergence, typically below 2 Å (Fig. 1). In contrast, off-target molecules were positioned further away and showed much greater variation. Notably, across the 16 benchmarks, the pose convergence metric generally outperformed the Bolt-2 binding-likelihood prediction (Fig. 1A, B; Table S3).

A Pose metric performance: Percentage of true ligands (“on-target” molecules) and false ligands (“off-target’ molecules) with pose convergence < 2.0 Å and distance from the reference binding site < 5.0 Å for each target. B Boltz-2 binding likelihood prediction: Percentage of ligand with predicted binding likelihood > 0.5 for both on-target and off-target ligands across 14 targets. C–J Scatter plots: Ligand pose convergence (RMSD) versus binding site distance for true on-target inhibitors (blue) and unrelated off-target compounds (gray), shown for ten representative targets: kinase CDK2 (C), protease thrombin (D), hydrolase PDE2 (E), kinesin KIF11/EG5 (F), phosphatase PTN11 (G), oxidoreductase DHFR (H), BCL2-like protein MCL1 (I), GPCR HCAR3 (J), lectin GAL3 (K), and PAS-domain EPAS1 (L). Scatter plots on the six remaining targets are provided in Fig. S1.
Specificity was especially strong for the kinases CDK2 and TYK2, the PAS-domain EPAS1, the hydrolase PDE2, the phosphatases PTP1B and PTN11, the oxidoreductase DHFR, the lectin GAL3, and the BCL-2-like protein MCL1. In contrast, the weakest specificity was observed for the GPCRs FFA2R and HCAR3, as well as the hydrolase BACE1, and to a lesser extent for the oxidoreductase COX-1, where some off-target molecules were still positioned near the binding site and exhibited low RMSD values. Similar trends were observed with the Boltz-2 model (Fig. S2).
Docking specificity in a molecular system with multiple binding sites
We next asked whether AI-based docking can also help identify the true binding site of a ligand in proteins that have multiple binding pockets. To address this, we selected Mycobacterium tuberculosis (Mtb) DNA gyrase as a model system, given its complex binding landscape and extensive characterization14,15. DNA gyrase contains several inhibitory sites that are targeted by chemically diverse compounds, making it an excellent case study for evaluating the potential of machine learning in drug discovery.
Fluoroquinolones (FQs) are the main class of inhibitors for this enzyme16,17,18, but inhibition can also be achieved by non-FQ compounds19,20, including novel bacterial type IIA topoisomerase inhibitors (NBTIs)21,22,23 and thiophene-based molecules24 (here referred as DNA gyrase allosteric inhibitors).
After validating the available crystal structures (Table S1), we predicted binding poses for several FQs lacking experimental structures, as well as a set of non-FQ compounds. These included ligands that bind DNA gyrase at sites distinct from the FQ pocket, inhibitors of benchmarks, and anti-tuberculous agents known to act on entirely different protein targets (Table S2).
On average, FQs clustered close to their known binding site, with a mean distance of less than 2 Å in models generated (Fig. 2B; Table S3). A similar pattern was observed for non-FQ inhibitors, such as those in the spiropyrimidinetrione class, represented by QPT-1 (QPT) and zoliflodacin (ZLF) (Fig. 2D). These compounds also function through the FQ mechanism by binding at FQ site19,20. NBTIs were consistently positioned within 2 Å of their known binding site, located approximately 10 Å from the FQ site and between the two scissile DNA bonds23,25 (Fig. 2B, E). This result was observed using both AF3 and Boltz-1, but not Boltz-2 (Table S3).

A Molecular system: Left panel shows the full DNA gyrase system used for calculation; right panel provides a close-up of the binding site region. The three known binding sites are highlighted: FQ site (green), non-catalytic (NC) site (blue), and allosteric site (pink). The gyrase is displayed as a gray molecular surface and DNA is shown in orange. B AF3 clustering: AF3 successfully separates FQs (green), NC-site binders or NBTIs (blue), and allosteric inhibitors (pink) from unrelated compounds (orange and gray) based on pose convergence and proximity to the FQ site. Similar results were obtained with Boltz-1 (Table S3). C Boltz-2 clustering: In contrast, Boltz-2 fails to achieve this separation, clustering all binders near the FQ site. D–F Docking poses generated by AF3: Close-up views for FQs (D), NBTIs (E), and allosteric inhibitors (F). D–F also presents a chart comparing AF3 pose convergence—defined as the percentage of true inhibitors and other ligands with pose convergence < 2.0 Å and a distance from the reference binding site < 5.0 Å—with Boltz-2 binding likelihood predictions, expressed as the percentage of inhibitors with a predicted binding likelihood > 0.5. A selected set of docked compounds is shown, including zoliflodacin (ZLF), trovafloxacin (TVF), AMK32b (ID8), gepotidacin (GPT), and thiophene 1 inhibitor (TH1). Each 3D image shows five representative AF3-predicted docking poses (i.e., the highest ipTM-scoring model for each seed) alongside the center of-mass of the reference FQ, MFX (gray sphere), superimposed on the reference crystal structure (PDB ID: 5BS8). Carbon atoms are color-coded as in (B), while oxygen and nitrogen atoms are shown in red and dark blue, respectively. The Mg2+ ions are represented as green spheres, the protein as a gray ribbon, and the DNA as an orange phosphate backbone with green–blue base sticks. Both the protein and DNA are shown in transparency. Protein visualizations were generated using PyMOL (The PyMOL Molecular Graphics System, Version 3.0, Schrödinger, LLC).
In contrast, regardless of the co-folding model used, thiophene-class allosteric inhibitors were rarely docked at their expected binding site, located approximately 30 Å from the FQ site (Fig. 2F)24. Instead, many of these inhibitors were mislocalized to the FQ or NC sites rather than their true allosteric site (Table S3). When docking was performed in the presence of gepotidacin (GPT) and sparfloxacin (SPF)—used to block the NC and FQ sites, respectively—the allosteric inhibitors remained confined to their correct binding site (Fig. 2F). Notably, poor prediction of allosteric binding site in the absence of occupied primary sites has recently been demonstrated using a dataset of 20 orthosteric/allosteric ligand pairs targeting 17 proteins26.
Finally, although off-target compounds and gyrase ATPase inhibitors adopted plausible binding poses according PoseBusters27 (Table S3), they were, on average, more distant from all three key binding sites of the molecular system under study (Fig. 2D).
Thus, the two criteria, pose convergence and proximity from the FQ site, effectively distinguished the three classes of gyrase inhibitors when using AF3 and Boltz-1 diffusion models. The clustering pattern clearly separated FQs, NBTIs, and allosteric inhibitors—for these latter inhibitors, particularly when the other two binding sites were already occupied by their respective ligands—from unrelated compounds, which were more broadly dispersed across the clusters (Fig. 2).
Competitive docking scoring and pairwise matrix
As shown in the previous section, diffusion-based co-folding models generate docking poses for any tested compounds, including those that do not specifically bind to the target protein. In the case of DNA gyrase, when two FQ molecules were docked simultaneously, only one occupied the catalytic FQ site, while the second often stacked against the DNA at the second catalytic site, which is only partially represented in the docking model used here. Furthermore, we previously showed that accurate docking of allosteric inhibitors on DNA gyrase can be achieved when both an FQ and an NBTI are included during docking inference (Fig. 2F). Building on these observations, we implemented a pairwise competitive docking approach to produce a scoring matrix, ranking compounds using a Competitive Docking Score (CDS) (Fig. 3).

Diffusion-based co-folding predictions were performed using a protein model bound to two competing ligands. The ligand that successfully occupies the active site was considered the winner of each competitive docking run. A Competitive Docking Score (CDS) was calculated from at least n independent runs per ligand pair. These scores were compiled into a pairwise matrix to rank ligands based on their cumulative CDS. Expressed as a percentage, the final CDS reflects the win rate across all pairwise docking runs. Protein visualizations were generated using PyMOL (The PyMOL Molecular Graphics System, Version 3.0, Schrödinger, LLC).
Competitive docking score correlation with inhibitor affinity
To assess the relevance of CDS rankings, we applied the method to benchmark proteins (Table S4) and compared the results with experimental affinity data reported in the literature28,29,30,31,32 or obtained from the Binding Database33 (Fig. 4).

A Rank concordance c-index between experimental inhibitory affinity and computed affinity or ranking, for AF3 pairwise docking strategy method (blue) and Boltz-2 binding affinity prediction (gray). Seventeen targets, containing nineteen binding sites were evaluated, with performance summarized by the rank concordance c-index. B–M Scatter plots show the relationship between the Competitive Docking Score (CDS) using AF3 and experimental pIC50 values for several systems studied. Each plot reports the Pearson correlation coefficient (r). An ordinary least squares regression line is added to illustrate the trend.
Using AF3, the rank concordance index (c-index)—a metric particularly relevant to this study, where correctly ordering compounds is more important than predicting their exact inhibitory values—showed strong agreement between CDS rankings and experimental affinities across many targets. Very strong and highly significance correlations were observed for lectin GAL3 (c = 0.89), kinase TYK2 (c = 0.87), protease thrombin (c = 0.86), kinase CDK2 (c = 0.78), GPCR HCAR3 (c = 0.77), kinesin KIF11 (c = 0.76), the DNA gyrase allosteric site (c = 0.76), phosphatases PTN11 and PTP1B (c = 0.75 and 0.74, respectively), and the DNA gyrase FQ site (c = 0.72). Strong correlations were obtained for GPCR FFA2R, PAS-domain EPAS1, BCL-2-like protein MCL1, hydrolase PDE2, and oxidoreductase DHFR (c = 0.66, 0.67, 0.68, 0.67, and 0.68, respectively). Moderate correlations were found for the DNA gyrase NC site (c = 0.62), hydrolase BACE1 (c = 0.62), and oxidoreductase COX-2 (c = 0.67), while COX-1 showed a weak correlation (c = 0.52) (Fig. S3; Table S4).
Importantly, the two systems with the weakest correlations (COX-1 and COX-2) also exhibited low docking pose convergence (Figs. 1 and 2), suggesting that even without competitive docking, inhibitors in these systems could not be reliably docked into the correct binding site.
It could be noted that pairwise competitive docking performed with Boltz-1/2 instead of AF3 generally produced weaker correlations with experimental IC50 values (Table S4), with no significant correlation observed for COX-1, COX-2, PDE2, PTP1B, and TYK2 systems.
We also found that the CDS rankings for COX-1 and COX-2 inhibitors were nearly identical (Pearson r = 0.96), indicating that AF3 could not reliably distinguish between the two cyclooxygenase isoenzymes (Table S5). This limitation is likely due to the high degree of conservation between their binding sites (Fig. S5). To further examine this issue, we tested four major DNA gyrase variants known to confer resistance to FQs by increasing IC50 or MIC values by several-fold30,34. In these cases, docking pose specificity remained largely unchanged, with only minor differences detected (Fig. S6). Similarly, the ranking of FQs using the pairwise competitive docking approach showed only minor shifts compared to the wild-type results (Fig. S6). These findings suggest that substituting one or two amino acids in the catalytic site is insufficient to significantly alter AF3’s binding predictions. This observation is consistent with recently published results showing that co-folding methods are unable to account for intentional perturbations in ligand-protein interaction modeling35.
Competitive docking approach vs. direct AI-based affinity prediction
Boltz-2 includes a machine-learning module for predicting protein-ligand affinity. We compared the performance of this tool with our competitive docking approach by assessing their correlations with experimental data (Fig. 4A). Overall, both methods demonstrated comparable performance across the 19 tested systems, with only minor differences. For instance, Boltz-2 performed slightly better for the BACE1, DHFR, EPAS1, FFA2R, PTP1B, and PDE2 systems, whereas competitive docking showed a slight advantage for the DNA gyrase allosteric site, CDK2, GAL3, HCAR3, PTN11, TYK2, and COX-2 systems.
DNA gyrase as a case study for evaluating CDS rankings
Using DNA-gyrase as a model, the CDS-based ranking method also demonstrates its ability to cluster inhibitors according to their inhibition rank. This capability was observed across all three protein sites. Specifically, ligands with higher CDS values generally corresponded to more potent inhibitors, while those with lower CDS values were typically weaker (Fig. 5A–C and S6; Tables S6–S8).

A 21 FQs on Mtb DNA-gyrase, B allosteric inhibitors, C NBTIs, and D 20 FQs on E. coli DNA gyrase. The bar charts show the percentage of inhibitors within each IC50 threshold category, based on their CDS ranking. IC50 thresholds are provided in Table S4. “n” indicates the number of inhibitors in each CDS category.
Because DNA gyrase is the primary FQ target in Escherichia coli36, we also measured IC50 values for the inhibition of E. coli growth by 22 FQs and compared them to their CDS rankings (Table S4 and Fig. S7A). Although the overall correlation was modest, a clear trend emerged: FQs with lower IC50 values generally ranked higher in the CDS list (Fig. S7B). Notably, since FQs are known to have limited aqueous solubility37, excluding two FQs predicted to be poorly soluble significantly improved the correlation (Fig. 5D). Importantly, CDS rankings could not be further refined by considering the secondary FQ target in E. coli, as the rankings generated using E. coli topoisomerase IV were nearly identical to those based on DNA gyrase (r = 0.99) (Figs. S7D, E).
All-at-Once docking strategy
To simplify the analysis and reduce computational cost, we evaluated AF3 docking performance by processing entire inhibitor classes in a single run, rather than relying on pairwise competitions. Using DNA gyrase as a model system, we tested three sets of inhibitors with AF3: 21 FQs, 24 NBTIs, and 12 allosteric inhibitors. For the allosteric set, docking was performed in the presence of GPT and SPF to block the other two competing binding sites.
Overall, the All-at-Once strategy yielded less detailed results compared to the pairwise competitive approach (Table S9). While some of the top-ranked compounds based on CDS values showed strong occupancy within the target binding site, several potent inhibitors were not identified among the leading competitors. Moreover, the number of distinct compounds effectively occupying the binding site was too limited for a reliable comparative analysis.
The All-at-Once strategy appears to be effective at identifying strong FQs from weaker FQs and non-FQ compounds. To evaluate its performance in a virtual screening context, we tested it on a compound library of 3155 FDA-approved molecules, including 46 FQs, representing 1.5% of the total library. The library was randomly divided into 124 sets containing 25–26 compounds each, and All-at-Once docking was performed using AF3 (Fig. 6A).

A Roadmap of the virtual screening performed on a library of 3155 compounds, including 46 FQs. B Percentage of FQs identified at different levels of the top-ranked compound list. Protein visualizations were generated using PyMOL (The PyMOL Molecular Graphics System, Version 3.0, Schrödinger, LLC).
This screening identified 147 top-ranking compounds, including 38 FQs, corresponding to a 25.9% enrichment. The eight FQs not selected as winners are known to have low inhibitory activity against Mtb DNA gyrase37 and mostly belong to the first-generation FQs (Table S10). Applying an additional filter based on pose convergence (cutoff of 2.5 Å) and proximity to the FQ binding site (cutoff of 2.0 Å), as described in Fig. 2, increased FQ enrichment to 77.8% among the 45 remaining compounds. Tightening these thresholds to 1.0 Å further boosted enrichment to 93.5%, with an enrichment factor of 62. These results closely matched the performance obtained with Boltz-2 for hit identification across the 3155-compound library (Fig. 6B).
Applying competitive docking to design more potent FQs
Given that competitive docking can assist in identifying the most effective compounds for a specific target, we investigated how this approach could be employed to design more potent FQs. As proof of concept, and without exhaustive exploration, we selected a set of 414 compounds from several thousand automatically generated using the STONED algorithm38. This selection focused on the chemical space surrounding the five top-ranked FQs identified by AF3 (Table S10). Each de novo compound was then evaluated in competitive docking against STF, the highest-ranked FQ, using the Mtb DNA gyrase model system.
Thirty-one of these newly designed compounds occupied the FQ binding site in at least 70% of the 100 generated models, suggesting a stronger binding potential than STF. Their Tanimoto structural similarity to STF ranged from 0.25 to 0.88 (Fig. S8). Since none of these compounds are listed in the CAS chemical database—indicating they have likely never been synthesized—we further filtered them based on predicted ADME properties. Ultimately, only eight de novo compounds exhibited favorable drug-likeness characteristics, solubility, and chemical synthetic accessibility.
link



