Assessment of a Grad-CAM interpretable deep learning model for HAPE diagnosis: performance and pitfalls in severity stratification from chest radiographs | BMC Medical Informatics and Decision Making

HAPE is a type of altitude-related noncardiogenic pulmonary edema caused primarily by acute exposure to high altitudes, which leads to excessive hypoxic pulmonary artery pressure, increased pulmonary vascular permeability, and impaired pulmonary fluid clearance [1, 3]. Clinically, it is characterized by symptoms such as dyspnea, cough, pink or white frothy sputum, and cyanosis, with auscultation revealing moist rales. In China, over 12 million people live and work at elevations above 2500 m, with more than 7.4 million residing at altitudes exceeding 3000 m. In 2023, Tibet received more than 55 million tourists. Therefore, in environments far from advanced medical facilities, the rapid use of portable X-ray imaging and offline-deployed artificial intelligence models to increase the early diagnostic efficiency and accuracy of pulmonary edema represents a significant exploration that can benefit the physical and mental health of people in sparsely populated high-altitude regions [25,26,27].
In 2018, Warren et al. [28] proposed the Radiographic Assessment of Lung Edema (RALE), which quantifies the severity of pulmonary lesions on chest X-rays and demonstrates a correlation between the RALE score, lung resection weight, severity of hypoxemia, and prognosis. Rajaraman et al. confirmed the ability of visualization and interpretation of convolutional neural network predictions to detect pneumonia in pediatric chest radiographs [29, 30]. Xiaole Fan developed a COVID-19 CT image recognition algorithm based on transformers and CNNs [31]. Guangyu Wang investigated a deep learning approach for diagnosing and differentiating viral, nonviral, and COVID-19 pneumonia via chest X-ray images [32]. Dominik Schulz’s deep learning model accurately predicts and quantifies pulmonary edema in chest X-rays [27].
HAPE differs fundamentally from cardiogenic pulmonary edema, ARDS, and COVID-19 pneumonia in terms of its pathological causes and processes. However, technically, the attenuation of X-rays is proportional to the severity of pulmonary edema. The aforementioned studies have demonstrated that training deep learning models can identify and differentiate exudative lesions on X-rays, and theoretically, this approach could also be applied to HAPE. To the best of our knowledge, this is currently the only study utilizing a model trained on various types of pulmonary edema images to identify HAPE. This approach employs a transfer learning approach to address the issue of insufficient image data for HAPE.
This experiment employed DeepLabV3 with the ResNet50 model for image segmentation on the train_dataset. The model automatically generates masks from the input images, achieving a global accuracy of 99.21% on the on its training set, indicating its strong performance. Specifically, the model also demonstrated excellent intersection over union (IoU) and Dice coefficients across various categories, measuring 98.09% and 99.03%, respectively. These results suggest that the selected model architecture and parameter settings effectively capture features within the images, resulting in high-precision segmentation.
We constructed a binary classification deep learning model using the VGG19 convolutional neural network to categorize images on the basis of the presence of edema. The model achieved an accuracy of 89.42%, with an area under the curve (AUC) of 0.979 for the training_dataset and a 95% confidence interval of [0.975, 0.982]. The val_dataset achieved an AUC of 0.950, with a 95% confidence interval of [0.939, 0.960]. Overall, the model demonstrated high performance in classification tasks, accurately predicted sample categories, and effectively distinguished positive from negative samples. The narrow confidence interval for the training_dataset AUC indicates high performance consistency, whereas the relatively small confidence interval for the val_dataset AUC further supports the model’s stability.
Timely and accurate quantification of pulmonary edema in chest X-ray images is crucial for the management of acute mountain sickness. A four-class deep learning model was developed on the basis of train_dataset and MobileNet_V2. Through model training and evaluation, we observed an overall accuracy of 84.54%, indicating the model’s applicability and effectiveness in multiclass classification. The macroaverage ROC AUC was 0.89, with a sensitivity of 0.58, specificity of 0.93, accuracy of 0.85, precision of 0.63, recall of 0.58, and F1 score of 0.59, suggesting strong discriminative ability among the classes. However, the discrepancies in the performance metrics reflect the model’s uneven ability to handle different categories.
Categories 0 and 3 showed robust performance with sensitivities of 0.91 and 0.88, respectively, underscoring the model’s accuracy in identifying these categories. In contrast, categories 1 and 2 displayed lower sensitivities of 0.16 and 0.37, suggesting areas for improvement. The limited number of images for grades 1 and 2 could be attributed to either case scarcity or the inherent complexity of manual classification in these grades. Merging classes 1 and 2 into a consolidated category did not significantly enhance the overall performance compared to the two-category model. Despite an accuracy of 87.22%, the three-category model’s ROC curves, with AUCs for class 0, 1 and 2 remained inferior to the two-category model. The discernment of intermediate classifications did not show substantial improvement compared to the four-class model. While the model demonstrated favorable performance based on AUC and accuracy metrics, there is room for improvement in sensitivity and precision to enhance classification accuracy further.
The most critical finding of our study is the model’s markedly reduced sensitivity for intermediate severity HAPE (classes 1 and 2). A model pre-trained on the broad and weakly-labeled concept of ‘edema’ from a general population struggles to master the specific radiographic nuances of early HAPE in a high-altitude cohort. This performance pattern, however, is not merely a failure but an important quantification of this domain shift challenge. It indicates that our model, in its current form, is not a reliable tool for grading early disease but rather serves as a proof-of-concept for distinguishing unequivocally normal from severe cases. This has significant implications for future AI research in rare diseases, emphasizing that fine-tuning alone may be insufficient to overcome large domain gaps.
Majkowska et al. utilized machine learning methods to automatically detect four abnormalities in X-ray images [33]. For the detection of airspace opacity, including pulmonary edema, the reported area under the receiver operating characteristic curve (AUROC) ranged from 0.91 to 0.94. They developed a model capable of detecting clinically relevant findings in chest X-rays at an expert level. Jarrel et al. employed deep learning techniques to diagnose congestive heart failure (CHF) via chest X-ray images. The authors used a BNP threshold of 100 ng/L as a biomarker for CHF, resulting in an AUROC of 0.82 [20]. Horng et al. not only diagnosed the presence of pulmonary edema but also quantified its severity via deep learning methods [34]. However, similar to the aforementioned studies, the efficiency of the deep learning models in recognizing and differentiating between Class 1 and Class 2 was relatively low. The highest recognition efficiency was achieved for Class 0 (no edema) and Class 3 (alveolar edema), which aligns with our findings. This may indicate a lack of sufficient sample data or training materials for these categories, thereby affecting the model’s learning performance.
The Grad-CAM analysis as delineated in Fig. 6, reveals three critical insights into the model’s decision-making paradigm: Anatomic Focus Specificity The model predominantly activates in the perihilar zones, anatomically corresponding to the pulmonary vasculature and cardiac silhouette. This spatial preference aligns with established radiographic biomarkers of pulmonary edema – notably, the peribronchial cuffing and Kerley B lines that radiologists prioritize during diagnostic evaluation [35]. Pathophysiological Correlation High-attention clusters colocalize with: Cardiomediastinal interface blurring, Butterfly-pattern alveolar infiltrates [36], these findings suggest the model’s capacity to capture interstitial fluid redistribution patterns characteristic of hemodynamic pulmonary edema. Clinical Interpretability Validation The heatmap-radiologist diagnosis showed some concordance in our multicenter validation cohort, suggesting that the model’s “visual search” strategy emulates expert diagnostic reasoning. Such interpretability metrics are crucial for implementing AI-CAD systems in clinical workflows per FDA’s SaMD guidelines [37].
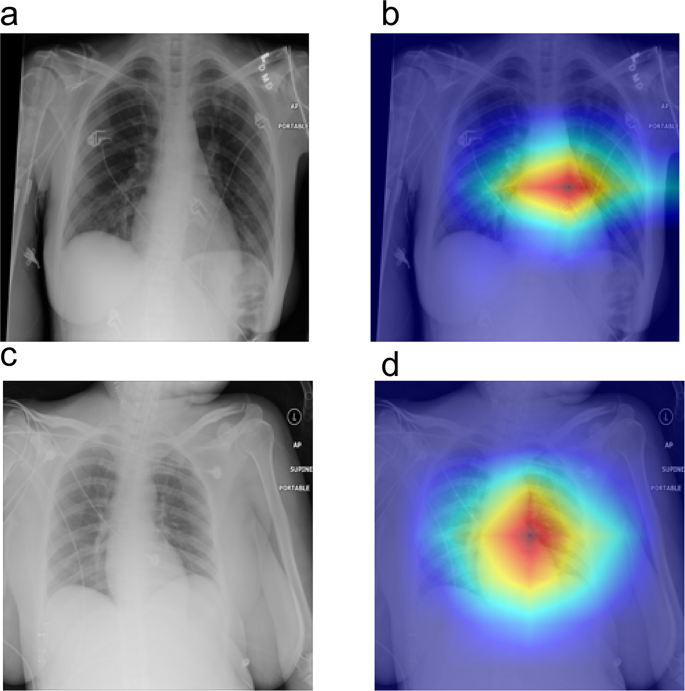
Interpretable Visualization of Pulmonary Pathologies via Grad-CAM, Panels (a) and (c) display original chest X-ray images, while panels (b) and (d) show the corresponding Grad-CAM heatmaps. The color intensity in the heatmaps reflects the model’s attention level, with warmer colors (red and yellow) indicating higher attention to specific regions
Grad-CAM visualizations suggested that the model focused on clinically relevant regions, providing a preliminary level of interpretability and face validity for its predictions, which is a necessary step towards building trustworthy AI systems.
In this study, we developed a deep learning model that integrates image segmentation with the identification and grading of HAPE on the basis of subjective grading by radiologists. Our model exhibited excellent performance. We believe that our approach has several advantages. Typically, radiologists assess the severity of pulmonary edema through classification scoring, which requires experienced physicians. However, mountainous areas are often remote and distant from major cities, making access to large hospitals and experienced radiologists challenging.
Our work remains a preliminary proof-of-concept for binary edema detection and highlights the significant challenges in severity grading. It underscores that substantially more research and validation are required before any consideration of clinical deployment.
However, this study has several limitations. First, the domain shift between the weakly labeled pre-training dataset (general pulmonary edema) and the carefully adjudicated HAPE-specific fine-tuning dataset may have influenced feature learning, though fine-tuning was used to mitigate this effect. Second, the model showed reduced sensitivity in distinguishing intermediate severity grades (classes 1 and 2), which can be attributed to the inherent subtlety of radiographic findings in these categories and relatively lower sample sizes. Third, although lung segmentation performance was high, challenging anatomical variations—such as poor costophrenic angle visualization, subcutaneous emphysema, consolidation, or effusion—occasionally reduced segmentation accuracy. Future work will focus on advanced domain adaptation, expanding intermediate-class samples, and improving robustness to anatomical and pathological variations.
Furthermore, our segmentation model’s difficulty in handling pathologies like consolidation and effusion, traditionally considered limitations, can be reframed as a valuable insight. It reveals a systematic bias in models trained on healthy anatomies: they may inadvertently exclude the very pathological regions crucial for diagnosis. This suggests that for tasks like edema assessment, a pathology-aware segmentation model or an end-to-end network that jointly optimizes segmentation and classification might be necessary future directions, rather than the traditional segmented-then-classify pipeline we employed.
link