Template Learning: Deep learning with domain randomization for particle picking in cryo-electron tomography
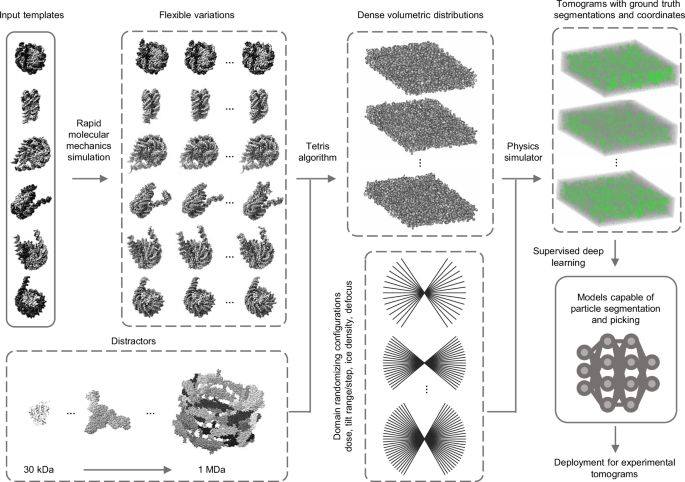
Simulations to train deep learning models on particle picking
Template Learning is a streamlined pipeline for simulating cryo-ET synthetic data for training deep learning models on particle picking. The fundamental goal of Template Learning is to train deep learning models in three aspects:
-
1.
Identifying targeted particles across diverse variations.
-
2.
Differentiating target particles from other structures, especially in crowded environments.
-
3.
Expanding the capabilities of the models to handle the inherent variations present in cryo-ET experimental data.
The practical application of these aspects is achieved through our proposed pipeline, illustrated in Fig. 1 and further elaborated in the subsequent sections.

Template structures are used as input, commonly differing by some structural variations of the target biomolecule (we mean by the target biomolecule, the structure that the deep learning model is desired to learn to annotate in cryo-ET tomograms, in the figure, it is the nucleosome). The input templates are augmented with flexible variations based on fast molecular mechanics simulations (see text for details). A general set of other proteins, different from the input templates, are used as “distractors” when generating synthetic data. The templates, along with their flexible variations and distractors are placed at random orientations in close proximity using the “Tetris algorithm”— an algorithm proposed in this work to enable fast simulation of high molecular crowding. A physics-based simulator (Parakeet28) is used to generate synthetic cryo-ET data using a set of different parameters that were chosen to produce domain randomization at the lowest cost, which are the electron dose, tilt range, tilt step, ice density, and defocus. The resulting output comprises tomograms, and ground truth segmentations, and coordinates corresponding to the templates. The generated data is subsequently utilized to train deep learning models (e.g., DeepFinder19) for experimental tomogram segmentation and particle picking. The PDB IDs used for templates in this figure are 2PYO, 7KBE, 7PEX, 7PEY, 7XZY, and 7Y00, and for distractors are 3QM1, 7NIU, and 6UP6. The display of structures was performed using ChimeraX54 and IMOD48 software.
To enable trained deep learning models to effectively identify various variations of the target biomolecule, we utilize following two strategies:
-
1.
We use multiple templates of one target biomolecule to account for compositional or significant conformational variabilities of the target particles.
-
2.
We generate random flexible variations of each template using Normal Mode Analysis (NMA—a method for fast molecular mechanics simulations, see Methods for details)39. The templates with their flexible variations are then simulated at uniformly random orientations to account for orientational variations.
Templates in this study are typically atomic structures. However, since the availability of atomic structures may vary for different target biomolecules, using lower resolution cryo-EM density maps (volumes) is also possible and is explored in a subsequent section.
The existing literature on domain randomization for object recognition in natural images simulates the target objects within a background containing a library of other random objects40. These additional objects, distractors, obscure or cast shadows on segments of the target objects. In this study, we selected 100 dissimilar protein assemblies, with diverse molecular weights ranging from 30 kDa to 1 MDa to serve as elements within the background (a comprehensive list of these proteins is available in Supplementary Fig. 1). We inspired our selection of these distractors from the work outlined in TomoTwin, knowing that these structures have different enough feature representations as interpreted by convolutional neural networks allowing to cluster them into corresponding classes. To achieve a similar effect of distractors for cryo-ET data, i.e., overlapping at the level of the tilt series and the missing wedge artifacts, these proteins must be positioned in close proximity to the templates during data modeling.
Recent works27,41 that involve simulating cryo-ET data for practical applications (e.g., deep learning) utilize an iterative brute-force random placement algorithm, involving the rotation of a duplicate of a molecule in each iteration to find a suitable non-overlapping position within the sample. Nonetheless, random placement leads to unstructured empty spaces between the molecules that prevent achieving highly dense samples.
Two other approaches simulate molecular crowding of cryo-ET data42,43. The first represents molecules as spheres, based on calculating the minimum bounding sphere for each molecule, and simulates crowding by optimizing a sphere-packing problem. However, this approach is limited to generating high crowding only for spherical-shaped molecules. The second builds on the first by including an additional step of molecular dynamics simulations to enhance packing density. However, routine application of this approach requires significant computational resources.
To achieve high molecular crowding while maintaining generality and efficiency, we drew inspiration for a simplified approach from a recent concept on packing generic 3D objects44, a solution we term the “Tetris algorithm.” Our algorithm, illustrated in Fig. 2, operates on an intuitive principle: it places molecules iteratively, with each iteration positioning a new molecule at a uniform random orientation as close as possible to those already placed (the user only needs to choose the minimum distance). For more details on the principles of the Tetris algorithm, see “Methods”.

A The flowchart of the algorithm, demonstrated in 2D for simplicity. The molecules that are desired to populate the sample are used as input. In the initial iteration, a molecule is positioned at the center of an expanded volumetric sample, forming the current output. In subsequent iterations, a new molecule undergoes binarization and dilation. By multiplying the binarized version with a large positive value and subtracting the result from the dilated version, an insertion shell is generated (referred to as ‘in-shell’). This insertion shell is then correlated with a binarized version of the current output volume, resulting in a correlation map (referred to as ‘c-map’). Positive values in the c-map (white voxels) represent viable positions for adding the new molecule without intersection, whereas negative and zero values (black and gray voxels, respectively) represent positions where intersection with existing molecules occurs or where the distance from other molecules is too large. If the correlation map is not entirely negative, the molecule is added to the current output at the index of the maximum entry, ensuring it avoids intersections while maintaining the highest compactness. B An example of the Tetris algorithm input and output at each iteration, alternating between placing templates (in green) and distractors (in red). The numbers in (B) correspond to the output at different iterations of the algorithm in (A). C An example of a Tetris algorithm output in 3D.
Simulations can partially replicate the domain of real-world data; however, variations in the real world can be unpredictable, and simulations simplify some complex phenomena for practical reasons. Thanks to domain randomization34,36, simulating an exact match of the real world is not necessary for training deep learning models capable of generalizing to real-world data. Instead, domain randomization focuses on training these models on diverse simulated scenarios to minimize the influence of the domain on their capabilities.
In this work, we employ a cryo-ET physics-based simulator called Parakeet28. Parakeet, and its MULTEM backend45, provide control over numerous parameters for simulating cryo-ET hardware and the behavior of biological samples during data recording. While many parameters related to the electron beam, lenses, detector, sample, and data acquisition strategy can be controlled, it might not be necessary to randomize every available parameter. On one hand, the combinations required to cover all conditions grow exponentially with the number of variables. On the other hand, some variables might have undesirable effects on the overall simulation. Here, we propose varying a few essential parameters that significantly influence the output simulation: the electron dose, defocus, tilting range, tilting step, and ice density, while giving other parameters default values. The electron dose is crucial for varying the SNR and the irradiation effects on the sample. The defocus plays a key role in controlling the Contrast Transfer Function (CTF)46. Varying the tilting range and step may not replicate experimental data acquisition strategies, however, it can expose the deep learning model to variations that increase its robustness, for example, to artifacts and imperfections resulting from tilt series alignment and reconstruction. Lastly, different ice densities can simulate the variability of sample thickness and solvent composition. A combination of three values for the defocus and two values for each of the other variables results in 48 combinations that we found sufficient for efficient domain randomization.
Benchmarking Template Learning for picking ribosomes in situ
In a recent study20, the first exhaustively annotated in situ cryo-ET dataset was published (EMPIAR-10988). Here, we use this dataset to evaluate the proposed Template Learning thoroughly.
In the following, we generate, based on the Template Learning workflow, a simulated dataset to train DeepFinder19 for ribosome annotation. We compare the performance of this model (i.e., DeepFinder trained solely on simulations) to the contemporary techniques, namely, DeePiCt20 and DeepFinder (trained solely on annotated experimental data), and 3D template matching (we will refer to it as template matching hereafter).
We organized our experiments based on two principles. First, we benchmark the complete method that integrates the aforementioned concepts of Template Learning. Then, we conducted experiments where specific elements from the data simulation pipeline were intentionally omitted (e.g., the structural variations of the target template, the crowding, etc.) to assess their impact on performance and to offer guidelines for future users. Second, acknowledging that the size of a simulated dataset is user-defined (generally in deep learning, the more training data, and the more diversity, the better), we ensured the feasibility of routine use of the method by limiting its time requirement to a maximum of two days on a single GPU (within one day on a typical computing node with 4 GPUs, see Code Availability for details).
To establish a Template Learning workflow, we proceeded with the following steps. We selected 6 eukaryotic ribosome PDB structures (PDB IDs: 4UG0, 4V6X, 6Y0G, 6Y2L, 6Z6M and 7CPU).
The 6 selected structures differ slightly in composition or conformation. By applying NMA, we generated 25 flexible variations for each structure (refer to the NMA section in Methods for details). While generating synthetic data, we employed the aforementioned general distractors (a full list of distractors is given in Supplementary Fig. 1). To simulate a crowded environment, we generated volumes from the template structures (i.e., the 6 ribosome PDBs and their flexible variations) and distractors with a sampling rate of 16 Å using Eman247 software e2pdb2mrc. We used the generated volumes as input to populate 48 crowded volumetric distributions of size 192 × 192 × 64 voxel3 using the Tetris algorithm, with ribosome templates appearing at a frequency of one for every 5 distractors to produce a balanced volumetric density ratio balance between distractors and templates in the simulated tomograms (see Tetris algorithm section in Methods for details). Subsequently, we fed the sample information (i.e., atomic structures of templates and distractors with the positions and orientations of crowding generated using the Tetris algorithm) to Parakeet28 to simulate 48 tilt series using a dose symmetric tilting scheme using the parameters listed in Supplementary Table 1. All other parameters adhered to default values for Volta Phase Plates (VPP) simulations within the Parakeet software. Subsequently, we binned the simulated tilt series to a similar sampling rate as the analyzed dataset (13 Å/pixel) and reconstructed them in IMOD48 using Weighted Back Projection. We used the tomograms and their corresponding ground-truth template coordinates (around 6500 simulated ribosomes) and segmentations (i.e., volumes of the same size as the tomograms where template instances appear in white over black background, illustrated in Supplementary Fig. 2) to train DeepFinder19 using its default parameter settings (the list of parameters is given in Supplementary Table 2).
We applied the DeepFinder simulations-trained model to the 10 VPP tomograms of the EMPIAR-10988 dataset without any preprocessing, resulting in segmentation maps, i.e., ribosome segmentations based on the decisions of the model. Subsequently, we employed the MeanShift algorithm offered by the DeepFinder software, using a clustering radius of 10 voxels, to extract both the coordinates and size of these segments (count of the number of voxels of each segment), with and without the application of a mask (specifically, a cytosol mask sourced from the dataset, utilized to eliminate false positives outside the mask; see Supplementary Note 1 for more details about these masks). We compare the extracted coordinates to the expert-validated annotations provided by the dataset at different levels of segment sizes used as a threshold (i.e., segments smaller than the threshold were removed). For this comparison, we maintained the criterion that two annotations—one from the output of the deep learning model and the other from the expert annotations—were considered to target the same particle if their spatial distance was within 10 voxels (a value consistent with the methodology of a previous study analyzed the same dataset20). An example of a tomogram, its corresponding segmentation map, and the coordinates set at a threshold that removed smaller objects compared to expert annotations are presented in Fig. 3. To evaluate the performance of the method, we computed the overall Recall and Precision with and without masking computed over all the tomograms jointly (refer to the Methods section for more details on the assessment metrics). The point where Precision and Recall curves intersect was utilized to determine the F1 score per tomogram for the dataset (shown in Fig. 4A). The results, further illustrated in Fig. 4I, J, reveal that Template Learning used to train DeepFinder exclusively on simulations, achieved state-of-the-art performance, outperforming template matching and previous deep learning methods trained solely on annotations derived from the same experimental dataset based on the median F1 score (compared to values reported in ref. 20).

Left: Display of a 2D central slice of a VPP tomogram. Middle: Segmentation of ribosome from the Template Learning workflow shown in black and white, with a 50% transparent overlay of the mask. Right: resulting annotations within the mask, obtained by converting segmentations to annotations using the MeanShift algorithm (provided by the DeepFinder19 software) using a clustering radius of 10 voxels, compared to expert annotations. Comparison includes True Positives (TP), False Positives (FP), and False Negatives (FN) relative to expert annotations. The tomogram, mask and expert annotations are sourced from the EMPIAR-10988 dataset.

Performance benchmarking and comparative analysis of the Template Learning on ribosome annotation on the S. pombe dataset (EMPIAR-10988, 10 VPP tomograms). A–H Performance measures for 8 variations of Template Learning settings: performance in all these experiments was evaluated on n = 10 independent tomograms from S. pombe in a single experiment, acknowledging that models trained solely on simulated data yield deterministic results on experimental data, making repeated identical evaluations unnecessary. The curves depict the overall Precision and Recall against the volume of the segmented region (horizontal axis). Precision is evaluated with and without masking (± mask). Boxplots show the F1 score per tomogram at the threshold where the Precision and Recall curves intersect, which often coincides with the highest overall F1 score (except for the curve shown in E (− mask), where the threshold was chosen based on the intersection point from the (+ mask) case). Box plot middle lines mark the mean value, and the edges indicate the 25th and 75th percentiles; whiskers indicate the range of existing data (no outliers were removed). I Table listing the differences between the experiments in (A–H), comparing their results based on their median F1 scores (bold value is the highest among proposed and previous techniques). J Table of previous results with their reported performance based on the findings in ref. 20. Table entries marked with an asterisk (*) indicate when values reported as the median of 3 independent cross-validation experiments, where each experiment used a random split of n = 8 tomograms for training and n = 2 tomograms for validation (out of the 10 original tomograms). Comparison of segmentations resulting from the different experiments on an example tomogram are shown in Supplementary Fig. 3. Descriptive statistics and statistical tests validating the significance of these Template Learning variations are provided in Supplementary Note 2.
To reflect on these results, it is essential to highlight that the previously reported findings for DeePiCt and DeepFinder were based on models trained with 8 out of 10 fully annotated tomograms, containing approximately 20,000 ribosome annotations20. Hence, previous evaluations were conducted on 2 out of 10 tomograms, for annotating the remaining 5000 ribosomes, utilizing three cross-validation splits (in each split, a random set of 8 tomograms was used for training and the remaining 2 tomograms for benchmarking). Additionally, each of the two models underwent a different training strategy to achieve optimal performance. In particular, in the case of DeepFinder, the training encompassed two classes—ribosomes and Fatty Acid Synthase (FAS)—as training solely on ribosomes led to suboptimal performance. On the other hand, DeePiCt was exclusively trained on ribosomes, as unlike DeepFinder, combining ribosomes and FAS during training led to suboptimal performance.
In contrast, the DeepFinder model trained solely on the simulations generated from the Template Learning workflow, not fine-tuned on any annotated experimental data, was benchmarked on the complete dataset (i.e., all 10 tomograms). Hence, despite the relatively modest increase in the F1 score for the Template Learning method (0.85, compared to the previous best of 0.83), it stands as a demonstration that deep learning can be trained effectively for picking a target structure, starting from only prior templates, and domain-randomized simulations. Also, a model of the same network architecture (DeepFinder) trained only on simulations outperforms its counterpart trained only on annotated experimental datasets for cryo-ET particle picking.
To systematically assess the contribution of each component of the Template Learning workflow, we performed a series of ablation and variation experiments, as detailed in the Supplementary Materials – Template Learning variations and Supplementary Notes 3 and 4. These analyses revealed that both incorporating multiple atomic structures and generating flexible molecular variations enhance annotation performance, and that introducing artificial flexible variations can partially compensate for the absence of multiple templates (Fig. 4A–C compared to Fig. 4D and Supplementary Fig. 3). The comparative impact of these design choices is illustrated by recall, precision, and F1 score. The inclusion of a diverse set of distractor molecules was found to be critical for high precision in particle picking; omitting or restricting distractor variability substantially increased false positive rates (Fig. 4E, F). Furthermore, simulating densely crowded environments via the Tetris algorithm (Supplementary Fig. 4) was essential, as reducing crowding in training simulations led to a marked drop in median F1 score (Fig. 4G). We also validated the use of low-resolution volumetric templates by developing a real-time conversion to pseudoatomic models compatible with physics-based simulations; this approach enabled training with reasonable recall, albeit at the expense of some precision (Fig. 4H). In addition, Template Learning could be readily adapted to different imaging domains, such as VPP and conventional defocus (DEF), and offered improved performance on cross-domain tasks compared to previous approaches (Fig. 5 and Supplementary Fig. 5).

Performance benchmarking and comparative analysis of Template Learning workflow applied to ribosome annotations within the S. pombe dataset (EMPIAR-10988, 10 DEF tomograms). A,B The curves depict the overall Precision and Recall against the volume of the segmented region (horizontal axis). Precision is evaluated with and without cytosol masking (±mask). Boxplots show the F1 score per tomogram at the threshold where the Precision and Recall curves intersect, which coincides with the highest overall F1 score. Performance in all these experiments was evaluated on n = 10 independent tomograms from S. pombe in a single experiment, acknowledging that models trained solely on simulated data yield deterministic results on experimental data, making repeated identical evaluations unnecessary. Box plot middle lines mark the mean value and the edges indicate the 25th and 75th percentiles; whiskers indicate the range of existing data (no outliers were removed). C On the top, a table listing the differences between the experiments in (A,B), comparing their results based on their median F1 scores. On the bottom, previously reported performance based on the findings in ref. 20.
Fine-tuning Template Learning-trained models using real data
The literature on domain randomization indicates that models pre-trained on simulations and subsequently fine-tuned on real-world data outperform models trained exclusively on real-world data35. This approach can be particularly valuable to cryo-ET particle picking studies that involve challenging-to-annotate molecules, or when training on domain randomization simulations alone does not yield satisfactory results.
In the same dataset used for benchmarking Template Learning on ribosomes in situ (EMPIAR-10988), another molecule, FAS, is annotated. Previous attempts based on template matching and supervised deep learning on experimental datasets have reported challenges in annotating FAS, attributed to its distinctive shell-like structure, where particles have lower SNR compared to ribosomes20.
In the following, we present the benchmarking of Template Learning combined with DeepFinder, initially trained exclusively on a simulated dataset, on FAS annotation in EMPIAR-10988 tomograms. Subsequently, we gradually introduce fine-tuning based on the experimental data annotations to assess whether performance can be improved.
To establish a Template Learning workflow for FAS, we utilized two PDB structures (PDB IDs: 4V59 and 6QL5). By using NMA, we generated 80 flexible variations for each structure (refer to the NMA section in the Methods for details). We kept the remaining parameters for simulated data generation consistent with those aforementioned for the ribosome study, except for training the models on sphere segmentations due to the shell-like structure of FAS.
The F1 score results of the simulation-trained DeepFinder model for FAS annotations are presented in Fig. 7. The results show that on the VPP dataset, the DeepFinder model trained on the Template Learning workflow outperformed that trained exclusively on experimental data. The results on the DEF dataset gave an F1 score ranging up to 22%, outperforming previous attempts that failed to pick any particle, based on the results reported in ref. 20. However, these results can still benefit from improvements compared to previously obtained results with DeePiCt trained on annotated experimental data.
We proceeded to fine-tune the simulations-trained DeepFinder model progressively using annotations from experimental data. In the following experiment, we fine-tuned it using annotations from 2 VPP tomograms containing approximately 150 FAS annotations. We kept all the DeepFinder training parameters to default (listed in Supplementary Table 2), except for reducing the number of steps and epochs to 10 steps for 10 epochs (in place of 100 steps for 100 epochs) to avoid overfitting (since the number of training data examples is low). We performed three cross-validation experiments (i.e., in each experiment, the data is randomly split into 2 tomograms for training and 8 for benchmarking). The results presented in Fig. 6 show a significant increase in the median F1 score compared to training on simulated data only. Notably, de Teresa-Trueba et al.20 previously reported that training a deep learning model with fewer than 600 particles was ineffective. Therefore, the ability to fine-tune our model with only 150 particles after pre-training on simulations represents a major advancement over previous supervised methods that required larger datasets to train models from scratch using only experimental annotations.

Performance benchmarking and comparative analysis of Template Learning applied to FAS annotation within the S. pombe dataset (EMPIAR-10988, 10 DEF tomograms). The results show the performance of the Template Learning workflow used for training DeepFinder on simulations, followed by different levels of fine-tuning on experimental data, compared to results based on the findings in ref. 20. Table entries marked with an asterisk (*) indicate when values reported as the median of 3 independent cross-validation experiments, where each experiment used a random split of either n = 2 or 8 tomograms for training and n = 8 or 2 tomograms for validation (out of the 10 original tomograms, indicated in the table). Otherwise (for entries without an asterisk), performance was assessed on all n = 10 tomograms in a single experiment, acknowledging that models trained solely on simulated data yield deterministic outcomes on experimental data, making repeated identical evaluations unnecessary. Box plot middle lines mark the mean value, and the edges indicate the 25th and 75th percentiles; whiskers indicate the range of existing data (no outliers were removed).
In a subsequent experiment, we fine-tuned the simulation-trained DeepFinder model on 8 VPP tomograms containing approximately 600 FAS annotations and inferred the model on the remaining 2 tomograms, using 3 cross-validation splits. Again, we kept all the DeepFinder training parameters to default, except for the number of steps and epochs, keeping 10 steps for 30 epochs (again, to prevent overfitting). The results presented in Fig. 6 show a significant increase in the median F1 score, outperforming previous supervised deep learning methods trained solely on experimental data.
Finally, we performed a cross-domain experiment, where we fine-tuned the simulation-trained DeepFinder model on the VPP dataset and applied this model to annotate FAS in the DEF dataset after SM preprocessing. Consistent with previous findings, the results presented in Fig. 6 show that training on simulations and fine-tuning on experimental data outperforms training only on experimental data.
Template Learning improves precision and isotropy in nucleosome picking
In cryo-ET data processing, localizing a target biomolecule in a new dataset is a common objective. The application of supervised deep learning to this task requires an initial training dataset of particles of interest extracted directly from the new data. Such a dataset is usually in the order of thousands of particles in different orientations, where more particles are needed for better performance. Manually annotating these particles is time-consuming, making template matching followed by manual elimination of obvious false positives and further curation through STA and classification the most common approach19,20. The principle of Template Learning offers a time and computing-efficient alternative to this complex procedure by annotating new data directly in a single step using a model trained on synthetic data.
In this section, we explore the efficiency of Template Learning for nucleosome annotation within a new cryo-ET dataset of partially decondensed mitotic chromosomes in vitro (refer to the Method section for sample preparation details) and compare it with a template matching-based annotation routine.
A tomogram central slice of our data is shown in Fig. 7A. Despite this dataset being isolated chromosomes in vitro, it contains an abundance of structures other than nucleosomes that can be false positives which are: DNA linkers, gold nanoparticles, percoll, and the non-histone components of chromatin.

A Central slice and zoomed-in 10-slice average of a cryo-ET tomogram of partially decondensed mitotic chromosomes in vitro. B,C The results of applying Template Learning (6 PDB structures as input, other parameters kept default) and traditional template matching (using PyTom with a nucleosome template) for annotating nucleosomes within the tomogram in (A), followed by reference-free STA in Relion V4. The experiments were repeated on 4 independent tomograms (N = 4) with similar results. B Top: score map resulting from training DeepFinder on nucleosome annotations using simulated datasets only from the proposed Template Learning workflow. Below: results of STA using 18 k annotations extracted using a 6 voxel clustering radius of the MeanShift algorithm (provided by the DeepFinder software). The results show initial model generation, followed by two stages of auto-refinement, and the angular distribution of averaged particles. The refinement has converged without particle curation, and the angular distribution of averaged particles is isotropic. C Top: score map resulting from traditional template matching. Below: results of STA using the top 18 k local maxima from the score map. The first step was an initial model generation, followed by a stage of refinement. The first stage of refinement did not improve the resolution of the initial model, therefore, it was followed by simultaneous classification and alignment into 10 classes. The particles corresponding to nucleosome-like classes (57% of the original 18 k particles) were used for two stages of refinement. The results show a slightly lower resolution average than the one obtained using Template Learning and show an anisotropic angular distribution.
To establish a Template Learning workflow, we utilized six nucleosome template structures with PDB IDs: 2PYO, 7KBE, 7PEX, 7PEY, 7XZY, and 7Y00.
By applying NMA, we generated 100 flexible variations for each structure (more details can be found in the NMA section of “Methods”). Notably, these templates represent different compositional variations of the nucleosome. The 7PEX structure includes the H1 protein, while the other five templates lack H1 but exhibit varying DNA linker lengths.
The remaining parameters of the Template Learning workflow were configured similarly to those used for the workflow established for ribosome and FAS annotation, with a few notable exceptions explained below.
Firstly, recognizing the smaller size of nucleosomes compared to ribosomes, we adjusted the frequency of appearance of nucleosome templates to one nucleosome for every two distractors (in contrast to one ribosome for every five distractors). This adjustment aimed to maintain a balanced volumetric density ratio between distractors and templates in the simulated tomograms.
Secondly, recognizing that nucleosomes have fewer atoms than ribosomes, we observed increased speed in the execution of the physics-based simulator (i.e., Parakeet). This allowed for the generation of larger tomograms for training, all within the same runtime for the ribosome study (see Code Availability for details). In particular, the tomogram size generated in the Tetris algorithm here was set to 256 x 256 x 64 (for a pixel size of 16 Å), in contrast to 192 x 192 x 64 used for ribosome simulations.
Lastly, we adjusted the pixel size of the simulated data to 8 Å through binning, closely approximating the pixel size of our experimental tomogram, which had been previously binned four times before annotation (the unbinned pixel size was 2.075 Å).
Consequently, we trained DeepFinder on the resulting Template Learning simulations and applied it to segment our data. The corresponding segmentation map (score map), is shown in Fig. 7B. Upon visual inspection, it is evident that that model has high Precision in localizing nucleosomes. Subsequently, we employed the MeanShift algorithm from the DeepFinder software to extract the corresponding annotations for the nucleosomes, utilizing a clustering radius of 6 pixels, approximately equivalent to the nucleosome’s radius at this pixel size. We excluded particles close to the edges of the tomogram, the air-water interface, and the sample-carbon interface.
Following this process, we obtained 18 k annotated particles and proceeded with the reference-free STA procedure using Relion V4.015 by generating an initial model from the data, followed by two stages of refinement at binning 4 and binning 2, resulting in resolving the nucleosome structure at 12.8 Å (at 0.143 FSC threshold) resolution shown in Fig. 7B.
The angular distribution of the averaged particles seems isotropic, showcasing that the Template Learning method is not biased to certain orientations.
To further verify the Precision of nucleosome identification, we classified the particles into 10 and 50 classes (presented in Supplementary Fig. 6). All class average results showed well-recognizable nucleosome shapes with DNA gyres, with the major differences being in the heterogeneity of the DNA entry-exit segments. We also performed a local resolution assessment using ResMap49 (presented in Supplementary Fig. 7) and observed that resolution has ranged from 12–20 Å, with the highest resolution in the DNA close to the core histones, and the lowest resolution around the DNA entry-exit segments.
Our averaging and classification results show that our Template Learning workflow generated precise nucleosome annotations of uniform orientations, facilitating high-throughput STA without the need for multiple rounds of classification to eliminate false-positive particles.
We applied template matching starting from a nucleosome template structure generated from the PDB 2PYO using the default procedure in PyTom18. We then applied the template matching procedure to our new data using an angular sampling of 7° increments (45,123 rotations).
The score map is shown in Fig. 7C. The visual inspection of the score map shows some high signal for some nucleosomes also identified by the Template Learning procedure (Fig. 7A), but also obvious false positive signal for other objects (an example of high false signal for the Percoll particles is shown at the center of the zoomed-in image in Fig. 7C).
To have a meaningful and fair comparison with the Template Learning procedure, we extracted from the same tomographic region the 18 k particles showing the highest cross-correlation scores. Unlike the results of Template Learning, a brief visual inspection of the template matching peaks showed 50 obvious false positives corresponding to the 10 nm gold and Percoll particles. The false positives were removed, and the data were processed Relion V4.015. The reference-free initial model was generated, followed by a stage of refinement at binning 4. Unlike Template Learning, the refinement did not result in a significant improvement of the resolution of the initial model (judged by the shape and the FSC), indicating the presence of a significant ratio of non-nucleosome particles among the annotations. In agreement with this assumption, classification, and alignment into 10 classes at the same binning resulted in only 2 nucleosome-like classes that sum to 10.25 k picks (57% of the original picks). The particles of these 2 classes were selected for a further refinement process at binning 4 reached Nyquist resolution, and the further refinement at binning 2 led to 12.9 Å resolution, which is comparable to that achieved with Template Learning.
Importantly, the angular distribution of nucleosomes annotated by template matching showed a significant imbalance towards side views compared to round top views (Fig. 7C). Our simulations (Supplementary Fig. 8) showed that due to the cylindrical shape of nucleosomes and the missing wedge problem of cryo-ET data, the constrained cross-correlation between a template and the particles is a function of the orientation, where side views show higher cross-correlation than top and oblique views. This problem results in extracting only side views of nucleosomes at an adequate Precision.
The orientation bias of template matching hampers its efficiency for particle detection and is prone to resolution loss in STA. In our case, the resolution of the nucleosome was not affected significantly, because its pseudo-symmetric cylindrical shape allowed the complete 360° orientation range of side views to have cross-correlation sufficient for detection (Fig. 7C). However, this constraint may pose a more significant challenge when annotating particles with asymmetric non-spherical shapes. To further validate the Template Learning pipeline in a crowded cellular environment, we compared its performance with supervised deep learning and template matching for nucleosome annotation within a tomogram of a vitreous section from Drosophila embryonic brain tissue8,50,51. Our STA results demonstrated that Template Learning outperformed the other two approaches in terms of precision and orientational isotropy (see Supplementary Note 5). Notably, template matching, a manual expert annotation, and a supervised model trained on this annotation exhibited significant orientational bias, favoring nucleosome side views. Our results demonstrate that Template Learning is capable of overcoming this limitation.
link